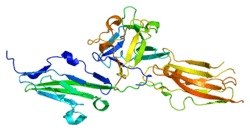Fibroblast growth factor receptor 2
Fibroblast growth factor receptor 2 (FGFR2) also known as CD332 (cluster of differentiation 332) is a protein that in humans is encoded by the FGFR2 gene residing on chromosome 10.[4][5] FGFR2 is a receptor for fibroblast growth factor.
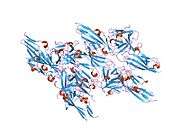
The protein encoded by this gene is a member of the fibroblast growth factor receptor family, where amino acid sequence is highly conserved between members and throughout evolution.[6] FGFR family members differ from one another in their ligand affinities and tissue distribution. A full-length representative protein consists of an extracellular region, composed of three immunoglobulin domains, a single hydrophobic membrane-spanning segment and a cytoplasmic tyrosine kinase domain. The extracellular portion of the protein interacts with fibroblast growth factors, setting in motion a cascade of downstream signals, ultimately influencing mitogenesis and differentiation. This particular family member is a high-affinity receptor for acidic, basic and/or keratinocyte growth factor, depending on the isoform.
Function
FGFR2 has important roles in embryonic development and tissue repair, especially bone and blood vessels. Like the other members of the fibroblast growth factor receptor family, these receptors signal by binding to their ligand and dimerisation (pairing of receptors), which causes the tyrosine kinase domains to initiate a cascade of intracellular signals. On a molecular level these signals mediate cell division, growth and differentiation.
Isoforms
FGFR2 has two naturally occurring isoforms, FGFR2IIIb and FGFR2IIIc, created by splicing of the third immunoglobulin-like domain. FGFR2IIIb is predominantly found in ectoderm derived tissues and endothelial organ lining, i.e. skin and internal organs.[7] FGFR2IIIc is found in mesenchyme, which includes craniofacial bone and for this reason the mutations of this gene and isoform are associated with craniosynostosis.
Interactions
Fibroblast growth factor receptor 2 has been shown to interact with FGF1.[8][9][10]
The spliced isoforms, however differ in binding:[11]
- FGFR2IIIb binds to FGF-1, -3, -7, -10, -22
- FGFR2IIIc binds to FGF-1, -2, -4, -6, -8, -9, -17 and -18
These differences in binding are not surprising, since FGF ligand is known to bind to the second and third immunoglobulin domain of the receptor.
Clinical significance
Mutations (changes) are associated with numerous medical conditions that include abnormal bone development (e.g. craniosynostosis syndromes) and cancer.
Craniosynostosis syndromes
- Apert syndrome, the best-known type of acrocephalosyndactyly, characterized by abnormalities of the skull and face, such as a cleft palate, and of the hands and feet.
- Antley-Bixler syndrome, characterized by trapezoidal, craniofacial and skeletal synostosis, plus camptodactyly), inherited as a recessive trait.
- Pfeiffer syndrome, another type of acrocephalosyndactyly, includes broad thumbs and large toes, inherited in autosomal dominant fashion.
- Crouzon syndrome, a craniofacial disorder with no hand or foot problems.[12] and potential cleft palate, inherited as a dominant trait.
- Jackson–Weiss syndrome
Cancer
- Breast cancer, a mutation or single nucleotide polymorphism (SNP) in intron 2 of the FGFR2 gene is associated with a higher breast cancer risk; however the risk is only mildly increased from about 10% lifetime breast cancer risk in the average woman in the industrialized world, to 12-14% risk in carriers of the SNP.[13]
Missense mutations of FGFR2 have been found in endometrial cancer and melanoma.[14]
As a drug target
AZD4547 is a tyrosine kinase inhibitor which targets FGFR1-3. It has demonstrated early evidence of efficacy in gastric cancer patients with high level FGFR2 amplification (Cancer Discovery 2016). FPA144 is a monoclonal antibody that binds to FGFR2b (a form of FGFR2) and preventing binding of certain FGFs. In 2014, a clinical trial began to treat gastric tumours that overexpress FGFR2b.[15] Another approach of FGFR2 targeting is use of allosteric inhibitors. Alofanib is a novel first-in-class allosteric small-molecular inhibitor of FGFR2. It binds to the extracellular domain of FGFR2 and has an inhibitory effect on FGF2-induced phosphorylation. Principal benefits of allosteric inhibitors are high selectivity and low toxicity [Tsimafeyeu et al. ESMO Asia 2016]. A phase Ib clinical study protocol has been selected for ECCO-AACR-EORTC-ESMO Workshop on Methods in Clinical Cancer Research, better known as the ‘Flims’ Workshop and clinical study of safety and preliminary efficacy of alofanib will be initiated at the beginning of 2017.
Mutations
FGFR2 mutations are associated with craniosynostosis syndromes, which are skull malformations caused by premature fusion of cranial sutures and other disease features according to the mutation itself. Analysis of chromosomal anomalies in patients led to the identification and confirmation of FGFR2 as a cleft lip and/or palate locus.[16] On a molecular level, mutations that affect FGFR2IIIc are associated with marked changes in osteoblast proliferation and differentiation.[17] Alteration in FGFR2 signalling is thought to underlie the craniosynostosis syndromes. To date, there are two mechanisms of altered FGFR2 signalling. The first is associated with constitutive activation of FGFR, where the FGFR2 receptor is always signalling, regardless of the amount of FGF ligand.[18] This mechanism is found in patients with Crouzon and Pfeiffer syndrome. The second, which is associated with Apert syndrome is a loss of specificity of the FGFR2 isoform, resulting in the receptor binding to FGFs that it does not normally bind.[19]
See also
References
- GRCh38: Ensembl release 89: ENSG00000066468 - Ensembl, May 2017
- "Human PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- "Mouse PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- Houssaint E, Blanquet PR, Champion-Arnaud P, Gesnel MC, Torriglia A, Courtois Y, Breathnach R (Oct 1990). "Related fibroblast growth factor receptor genes exist in the human genome". Proceedings of the National Academy of Sciences of the United States of America. 87 (20): 8180–4. doi:10.1073/pnas.87.20.8180. PMC 54916. PMID 2172978.
- Dionne CA, Crumley G, Bellot F, Kaplow JM, Searfoss G, Ruta M, Burgess WH, Jaye M, Schlessinger J (Sep 1990). "Cloning and expression of two distinct high-affinity receptors cross-reacting with acidic and basic fibroblast growth factors". The EMBO Journal. 9 (9): 2685–92. doi:10.1002/j.1460-2075.1990.tb07454.x. PMC 551973. PMID 1697263.
- "Entrez Gene: FGFR2 fibroblast growth factor receptor 2 (bacteria-expressed kinase, keratinocyte growth factor receptor, craniofacial dysostosis 1, Crouzon syndrome, Pfeiffer syndrome, Jackson–Weiss syndrome)".
- Orr-Urtreger A, Bedford MT, Burakova T, Arman E, Zimmer Y, Yayon A, Givol D, Lonai P (Aug 1993). "Developmental localization of the splicing alternatives of fibroblast growth factor receptor-2 (FGFR2)". Developmental Biology. 158 (2): 475–86. doi:10.1006/dbio.1993.1205. PMID 8393815.
- Stauber DJ, DiGabriele AD, Hendrickson WA (Jan 2000). "Structural interactions of fibroblast growth factor receptor with its ligands". Proceedings of the National Academy of Sciences of the United States of America. 97 (1): 49–54. doi:10.1073/pnas.97.1.49. PMC 26614. PMID 10618369.
- Pellegrini L, Burke DF, von Delft F, Mulloy B, Blundell TL (Oct 2000). "Crystal structure of fibroblast growth factor receptor ectodomain bound to ligand and heparin". Nature. 407 (6807): 1029–34. doi:10.1038/35039551. PMID 11069186.
- Santos-Ocampo S, Colvin JS, Chellaiah A, Ornitz DM (Jan 1996). "Expression and biological activity of mouse fibroblast growth factor-9". The Journal of Biological Chemistry. 271 (3): 1726–31. doi:10.1074/jbc.271.3.1726. PMID 8576175.
- Ornitz DM, Xu J, Colvin JS, McEwen DG, MacArthur CA, Coulier F, Gao G, Goldfarb M (Jun 1996). "Receptor specificity of the fibroblast growth factor family". The Journal of Biological Chemistry. 271 (25): 15292–7. doi:10.1074/jbc.271.25.15292. PMID 8663044.
- Sagong B, Jung da J, Baek JI, Kim MA, Lee J, Lee SH, Kim UK, Lee KY (2014). "Identification of causative mutation in a Korean family with Crouzon syndrome using whole exome sequencing". Annals of Clinical and Laboratory Science. 44 (4): 476–83. PMID 25361936.
- Hunter DJ, Kraft P, Jacobs KB, Cox DG, Yeager M, Hankinson SE, Wacholder S, Wang Z, Welch R, Hutchinson A, Wang J, Yu K, Chatterjee N, Orr N, Willett WC, Colditz GA, Ziegler RG, Berg CD, Buys SS, McCarty CA, Feigelson HS, Calle EE, Thun MJ, Hayes RB, Tucker M, Gerhard DS, Fraumeni JF, Hoover RN, Thomas G, Chanock SJ (Jul 2007). "A genome-wide association study identifies alleles in FGFR2 associated with risk of sporadic postmenopausal breast cancer". Nature Genetics. 39 (7): 870–4. doi:10.1038/ng2075. PMC 3493132. PMID 17529973.
- Katoh M, Nakagama H (Mar 2014). "FGF receptors: cancer biology and therapeutics". Medicinal Research Reviews. 34 (2): 280–300. doi:10.1002/med.21288. PMID 23696246.
- Open-Label, Dose-Finding Study Evaluating Safety and PK of FPA144 in Patients With Advanced Solid Tumors
- Dixon MJ, Marazita ML, Beaty TH, Murray JC (2011). "Cleft lip and palate: understanding genetic and environmental influences". Nature Review Genetics (12): 167-178.
- Lee KM, Santos-Ruiz L, Ferretti P (Mar 2010). "A single-point mutation in FGFR2 affects cell cycle and Tgfbeta signalling in osteoblasts" (PDF). Biochimica et Biophysica Acta (BBA) - Molecular Basis of Disease. 1802 (3): 347–55. doi:10.1016/j.bbadis.2009.11.006. PMID 20004243.
- Webster MK, Donoghue DJ (Oct 1997). "Enhanced signaling and morphological transformation by a membrane-localized derivative of the fibroblast growth factor receptor 3 kinase domain". Molecular and Cellular Biology. 17 (10): 5739–47. doi:10.1128/mcb.17.10.5739. PMC 232422. PMID 9315632.
- Hajihosseini MK, Duarte R, Pegrum J, Donjacour A, Lana-Elola E, Rice DP, Sharpe J, Dickson C (Feb 2009). "Evidence that Fgf10 contributes to the skeletal and visceral defects of an Apert syndrome mouse model". Developmental Dynamics. 238 (2): 376–85. doi:10.1002/dvdy.21648. PMID 18773495.
Further reading
- McKeehan WL, Kan M (Sep 1994). "Heparan sulfate fibroblast growth factor receptor complex: structure-function relationships". Molecular Reproduction and Development. 39 (1): 69–81, discussion 81–2. doi:10.1002/mrd.1080390112. PMID 7999363.
- Johnson DE, Williams LT (1993). "Structural and functional diversity in the FGF receptor multigene family". Advances in Cancer Research. 60: 1–41. doi:10.1016/S0065-230X(08)60821-0. ISBN 978-0-12-006660-5. PMID 8417497. Cite journal requires
|journal=(help) - Park WJ, Meyers GA, Li X, Theda C, Day D, Orlow SJ, Jones MC, Jabs EW (Jul 1995). "Novel FGFR2 mutations in Crouzon and Jackson–Weiss syndromes show allelic heterogeneity and phenotypic variability". Human Molecular Genetics. 4 (7): 1229–33. doi:10.1093/hmg/4.7.1229. PMID 8528214.
- Marie PJ, Debiais F, Haÿ E (2003). "Regulation of human cranial osteoblast phenotype by FGF-2, FGFR-2 and BMP-2 signaling". Histology and Histopathology. 17 (3): 877–85. doi:10.14670/HH-17.877. PMID 12168799.
- Ibrahimi OA, Chiu ES, McCarthy JG, Mohammadi M (Jan 2005). "Understanding the molecular basis of Apert syndrome". Plastic and Reconstructive Surgery. 115 (1): 264–70. doi:10.1097/01.PRS.0000146703.08958.95 (inactive 2020-05-25). PMID 15622262.
External links
- GeneReviews/NIH/NCBI/UW entry on FGFR-Related Craniosynostosis Syndromes
- Fibroblast+Growth+Factor+Receptor+2 at the US National Library of Medicine Medical Subject Headings (MeSH)
- FGFR2 human gene location in the UCSC Genome Browser.
- FGFR2 human gene details in the UCSC Genome Browser.