Lysozyme
Lysozyme, also known as muramidase or N-acetylmuramide glycanhydrolase, is an antimicrobial enzyme produced by animals that forms part of the innate immune system. Lysozyme is a glycoside hydrolase that catalyzes the hydrolysis of 1,4-beta-linkages between N-acetylmuramic acid and N-acetyl-D-glucosamine residues in peptidoglycan, which is the major component of gram-positive bacterial cell wall.[1] This hydrolysis in turn compromises the integrity of bacterial cell walls causing lysis of the bacteria.
| Lysozyme | |||||||||
|---|---|---|---|---|---|---|---|---|---|
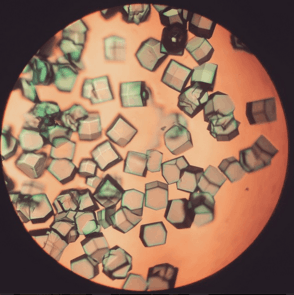 Lysozyme crystals stained with methylene blue. | |||||||||
| Identifiers | |||||||||
| EC number | 3.2.1.17 | ||||||||
| CAS number | 9001-63-2 | ||||||||
| Databases | |||||||||
| IntEnz | IntEnz view | ||||||||
| BRENDA | BRENDA entry | ||||||||
| ExPASy | NiceZyme view | ||||||||
| KEGG | KEGG entry | ||||||||
| MetaCyc | metabolic pathway | ||||||||
| PRIAM | profile | ||||||||
| PDB structures | RCSB PDB PDBe PDBsum | ||||||||
| Gene Ontology | AmiGO / QuickGO | ||||||||
| |||||||||
Lysozyme is abundant in secretions including tears, saliva, human milk, and mucus. It is also present in cytoplasmic granules of the macrophages and the polymorphonuclear neutrophils (PMNs). Large amounts of lysozyme can be found in egg white. C-type lysozymes are closely related to alpha-lactalbumin in sequence and structure, making them part of the same glycoside hydrolase family 22.[2] In humans, the C-type lysozyme enzyme is encoded by the LYZ gene.[3][4]
Hen egg white lysozyme is thermally stable, with a melting point reaching up to 72 °C at pH 5.0.[5] However, lysozyme in human milk loses activity very quickly at that temperature.[6] Hen egg white lysozyme maintains its activity in a large range of pH (6-9).[7] Its isoelectric point is 11.35.[8] The isoelectric point of human milk lysozyme is 10.5-11.[9]
Function and mechanism
The enzyme functions by attacking, hydrolyzing, and breaking glycosidic bonds in peptidoglycans. The enzyme can also break glycosidic bonds in chitin, although not as effectively as true chitinases.[10]
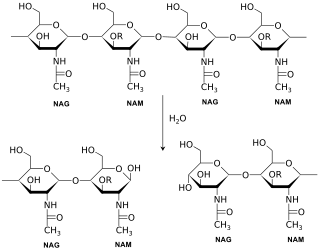
Lysozymes active site binds the peptidoglycan molecule in the prominent cleft between its two domains. It attacks peptidoglycans (found in the cell walls of bacteria, especially Gram-positive bacteria), its natural substrate, between N-acetylmuramic acid (NAM) and the fourth carbon atom of N-acetylglucosamine (NAG).
Shorter saccharides like tetrasaccharide have also shown to be viable substrates but via an intermediate with a longer chain.[11] Chitin has also been shown to be a viable lysozyme substrate. Artificial substrates have also been developed and used in lysozyme.[12]
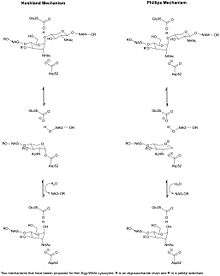
Mechanism
Phillips
The Phillips Mechanism proposed that the enzyme's catalytic power came from both steric strain on the bound substrate and electrostatic stabilization of an oxo-carbenium intermediate. From X-ray crystallographic data, Phillips proposed the active site of the enzyme, where a hexasaccharide binds. The lysozyme distorts the fourth sugar (in the D or -1 subsite) in the hexasaccharide into a half-chair conformation. In this stressed state, the glycosidic bond is more easily broken.[13] An ionic intermediate containing an oxo-carbenium is created as a result of the glycosidic bond breaking.[14] Thus distortion causing the substrate molecule to adopt a strained conformation similar to that of the transition state will lower the energy barrier of the reaction.[15]
The proposed oxo-carbonium intermediate was speculated to be electrostatically stabilized by aspartate and glutamate residues in the active site by Arieh Warshel in 1978. The electrostatic stabilization argument was based on comparison to bulk water, the reorientation of water dipoles can cancel out the stabilizing energy of charge interaction. In Warshel's model, the enzyme acts as a super-solvent, which fixes the orientation of ion pairs and provides super-solvation (very good stabilization of ion pairs), and especially lower the energy when to ions are close to each other.[16]
The rate-determining step(RDS) in this mechanism is related to formation of the oxo-carbenium intermediate. There were some contradictory results to indicate the exact RDS. By tracing the formation of product (p-nitrophenol), it was discovered that the RDS can change over different temperatures, which was a reason for those contradictory results. At a higher temperature the RDS is formation of glycosyl enzyme intermediate and at a lower temperature the break down of that intermediate.[17]
Koshland

In an early debate in 1969, Dahlquist proposed a covalent mechanism for lysozyme based on kinetic isotope effect,[19] but for a long time the ionic mechanism was more accepted. In 2001, a revised mechanism was proposed by Vocadlo via a covalent but not ionic intermediate. Evidence from ESI-MS analysis indicated a covalent intermediate. A 2-fluoro substituted substrate was used to lower the reaction rate and accumulate an intermediate for characterization.[20] The amino acid side-chains glutamic acid 35 (Glu35) and aspartate 52 (Asp52) have been found to be critical to the activity of this enzyme. Glu35 acts as a proton donor to the glycosidic bond, cleaving the C-O bond in the substrate, whereas Asp52 acts as a nucleophile to generate a glycosyl enzyme intermediate. The Glu35 reacts with water to form hydroxyl ion, a stronger nucleophile than water, which then attacks the glycosyl enzyme intermediate, to give the product of hydrolysis and leaving the enzyme unchanged.[21] This covalent mechanism was named after Koshland, who first proposed this type of mechanism.[22]
More recently, quantum mechanics/ molecular mechanics (QM/MM) molecular dynamics simulations have been using the crystal of HEWL and predict the existence of a covalent intermediate.[23] Evidence for the ESI-MS and X-ray structures indicate the existence of covalent intermediate, but primarily rely on using a less active mutant or non-native substrate. Thus, QM/MM molecular dynamics provides the unique ability to directly investigate the mechanism of wild-type HEWL and native substrate. The calculations revealed that the covalent intermediate from the Koshland mechanism is ~30 kcal/mol more stable than the ionic intermediate from the Phillips mechanism.[23] These calculation demonstrate that the ionic intermediate is extremely energetically unfavorable and the covalent intermediates observed from experiments using less active mutant or non-native substrates provide useful insight into the mechanism of wild-type HEWL.
Inhibition
Imidazole derivatives can form a charge-transfer complex with some residues (in or outside active center) to achieve a competitive inhibition of lysozyme.[24] In Gram-negative bacteria, the lipopolysaccharide acts as a non-competitive inhibitior by highly-favored binding with lysozyme.[25]
Non-enzymatic action
Despite that the muramidase activity of lysozyme has been supposed to play the key role for its antibacterial properties, evidence of its non-enzymatic action was also reported. For example, blocking the catalytic activity of lysozyme by mutation of critical amino acid in the active site (52-Asp -> 52-Ser) does not eliminate its antimicrobial activity.[26] The lectin-like ability of lysozyme to recognize bacterial carbohydrate antigen without lytic activity was reported for tetrasaccharide related to lipopolysaccharide of Klebsiella pneumoniae.[27] Also, lysozyme interacts with antibodies and T-cell receptors.[28]
Enzyme conformation changes
Lysozyme exhibits two conformations: an open active state and a closed inactive state. The catalytic relevance was examined with single walled carbon nanotubes (SWCN) field effect transistors (FETs), where a singular lysozyme was bound to the SWCN FET.[29] Electronically monitoring the lysozyme showed two conformations, an open active site and a closed inactive site. In its active state lysozyme is able to processively hydrolyze its substrate, breaking on average 100 bonds at a rate of 15 per second. In order to bind a new substrate and move from the closed inactive state to the open active state requires two conformation step changes, while inactivation requires one step.
Role in disease and therapy
| lysozyme family | |||||||
|---|---|---|---|---|---|---|---|
| Identifiers | |||||||
| Aliases | Glyco_hydro_22_lysIPR000974lysozymeLysozyme1,4-N-AcetylmuramidaseDelvozymeGlobulin G1Lydium-KLPMucopeptide glucohydrolaseMuramidaseN,O-DiacetylmuramidasePeptidoglycan N-acetylmuramoylhydrolasemucopeptide N-acetylmuramoylhydrolase | ||||||
| External IDs | GeneCards: | ||||||
| Orthologs | |||||||
| Species | Human | Mouse | |||||
| Entrez |
|
| |||||
| Ensembl |
|
| |||||
| UniProt |
|
| |||||
| RefSeq (mRNA) |
|
| |||||
| RefSeq (protein) |
|
| |||||
| Location (UCSC) | n/a | n/a | |||||
| PubMed search | n/a | n/a | |||||
| Wikidata | |||||||
| |||||||
Lysozyme is part of the innate immune system. Reduced lysozyme levels have been associated with bronchopulmonary dysplasia in newborns.[30] Piglets fed with human lysozyme milk can recover from diarrheal disease caused by E. coli faster. The concentration of lysozyme in human milk is 1,600 to 3,000 times greater than the concentration in livestock milk. Human lysozyme is more active than hen egg white lysozyme. A transgenic line of goats (with a founder named "Artemis") were developed to produce milk with human lysozyme to protect children from diarrhea if they can't get the benefits of human breastfeeding.[31][32]
Since lysozyme is a natural form of protection from Gram-positive pathogens like Bacillus and Streptococcus,[33] it plays an important role in immunology of infants in human milk feeding.[34] Whereas the skin is a protective barrier due to its dryness and acidity, the conjunctiva (membrane covering the eye) is, instead, protected by secreted enzymes, mainly lysozyme and defensin. However, when these protective barriers fail, conjunctivitis results.
In certain cancers (especially myelomonocytic leukemia) excessive production of lysozyme by cancer cells can lead to toxic levels of lysozyme in the blood. High lysozyme blood levels can lead to kidney failure and low blood potassium, conditions that may improve or resolve with treatment of the primary malignancy.
Serum lysozyme is much less specific for diagnosis of sarcoidosis than serum angiotensin converting enzyme; however, since it is more sensitive, it is used as a marker of sarcoidosis disease activity and is suitable for disease monitoring in proven cases.[35]
Chemical synthesis
The first chemical synthesis of a lysozyme protein was attempted by Prof. George W. Kenner and his group at the University of Liverpool in England.[36] This was finally achieved in 2007 by Thomas Durek in Steve Kent's lab at the University of Chicago who made a synthetic functional lysozyme molecule.[37]
Other applications
Lysozyme crystals have been used to grow other functional materials for catalysis and biomedical applications.[38][39][40] Lysozyme is a commonly used enzyme for lysing gram positive bacteria.[41] Due to the unique function of lysozyme in which it can digest the cell wall and causes osmotic shock (burst the cell by suddenly changing solute concentration around the cell and thus the osmotic pressure), lysozyme is commonly used in lab setting to release proteins from bacterium periplasm while the inner membrane remains sealed as vesicles called the spheroplast.[42][43]
For example, E. coli can be lysed using lysozyme to free the contents of the periplasmic space. It is especially useful in lab setting for trying to collect the contents of the periplasm.[1] Lysozyme treatment is optimal at particular temperatures, pH ranges, and salt concentrations. Lysozyme activity increases with increasing temperatures, up to 60 degrees Celsius, with a pH range of 6.0-7.0. The salts present also affect lysozyme treatment, where some assert inhibitory effects, and others promote lysis via lysozyme treatment. Sodium chloride induces lysis, but at high concentrations, it is an active inhibitor of lysis. Similar observations have been seen with the use of potassium salts. Slight variations are present due to differences in bacterial strains.[44]
History
The antibacterial property of hen egg white, due to the lysozyme it contains, was first observed by Laschtschenko in 1909.[45] The bacteria-killing activity of nasal mucous was demonstrated in 1922 by Alexander Fleming, the discoverer of penicillin, who coined the term lysozyme.[46] Fleming reported, saying. "As this substance has properties akin to those of ferments I have called it a 'Lysozyme'."[47] Fleming went on to show that an enzymic substance was present in a wide variety of secretions and was capable of rapidly lysing (i.e. dissolving) different bacteria, particularly a yellow "coccus" that he studied".[48]
Lysozyme was first crystallised by Edward Abraham in 1937, enabling the three-dimensional structure of hen egg white lysozyme to be described by David Chilton Phillips in 1965, when he obtained the first 2-ångström (200 pm) resolution model via X-ray crystallography.[49][50] The structure was publicly presented at a Royal Institution lecture in 1965.[51] Lysozyme was the second protein structure and the first enzyme structure to be solved via X-ray diffraction methods, and the first enzyme to be fully sequenced that contains all twenty common amino acids.[52] As a result of Phillips' elucidation of the structure of lysozyme, it was also the first enzyme to have a detailed, specific mechanism suggested for its method of catalytic action.[53][54][55] This work led Phillips to provide an explanation for how enzymes speed up a chemical reaction in terms of its physical structures. The original mechanism proposed by Phillips was more recently revised.[20]
See also
References
- Manchenko GP (1994). "Lysozyme". Handbook of Detection of Enzymes on Electrophoretic Gels. Boca Raton, Fla.: CRC Press. p. 223. ISBN 978-0-8493-8935-1.
- Williams S, Vocadlo D. "Glycoside hydrolase family 22". Cazypedia. Retrieved 11 April 2017.
- Yoshimura K, Toibana A, Nakahama K (January 1988). "Human lysozyme: sequencing of a cDNA, and expression and secretion by Saccharomyces cerevisiae". Biochemical and Biophysical Research Communications. 150 (2): 794–801. doi:10.1016/0006-291X(88)90461-5. PMID 2829884.
- Peters CW, Kruse U, Pollwein R, Grzeschik KH, Sippel AE (July 1989). "The human lysozyme gene. Sequence organization and chromosomal localization". European Journal of Biochemistry. 182 (3): 507–16. doi:10.1111/j.1432-1033.1989.tb14857.x. PMID 2546758.
- Venkataramani S, Truntzer J, Coleman DR (April 2013). "Thermal stability of high concentration lysozyme across varying pH: A Fourier Transform Infrared study". Journal of Pharmacy & Bioallied Sciences. 5 (2): 148–53. doi:10.4103/0975-7406.111821. PMC 3697194. PMID 23833521.
- Chandan RC, Shahani KM, Holly RG (October 1964). "Lysozyme Content of Human Milk". Nature. 204 (4953): 76–7. Bibcode:1964Natur.204...76C. doi:10.1038/204076a0. PMID 14240122.
- "Lysozyme, Product information" (PDF). Sigma-Aldrich.
- "Lysozyme, Product information" (PDF). Sigma-Aldrich.
- Parry Jr. RM, Chandan RC, Shahani KM (1969). "Isolation and characterization of human milk lysozyme". Arch Biochem Biophysics. 103 (1): 59–65. doi:10.1016/0003-9861(69)90009-5. PMID 5778672.CS1 maint: uses authors parameter (link)
- Skujiņś J, Puķite A, McLaren AD (December 1973). "Adsorption and reactions of chitinase and lysozyme on chitin". Molecular and Cellular Biochemistry. 2 (2): 221–8. doi:10.1007/BF01795475. PMID 4359167.
- Sharon N (April 1967). "The chemical structure of lysozyme substrates and their cleavage by the enzyme". Proceedings of the Royal Society of London. Series B, Biological Sciences. 167 (1009): 402–15. Bibcode:1967RSPSB.167..402S. doi:10.1098/rspb.1967.0037. PMID 4382803.
- Höltje JV (1 January 1996). "Lysozyme substrates". Exs. Experientia Supplementum. 75: 105–10. doi:10.1007/978-3-0348-9225-4_7. ISBN 978-3-0348-9952-9. PMID 8765297.
- Blake CC, Johnson LN, Mair GA, North AC, Phillips DC, Sarma VR (April 1967). "Crystallographic studies of the activity of hen egg-white lysozyme". Proceedings of the Royal Society of London. Series B, Biological Sciences. 167 (1009): 378–88. Bibcode:1967RSPSB.167..378B. doi:10.1098/rspb.1967.0035. PMID 4382801.
- Dahlquist FW, Rand-Meir T, Raftery MA (October 1969). "Application of secondary alpha-deuterium kinetic isotope effects to studies of enzyme catalysis. Glycoside hydrolysis by lysozyme and beta-glucosidase". Biochemistry. 8 (10): 4214–21. doi:10.1021/bi00838a045. PMID 5388150.
- McKenzie HA, White FH (1991). "Lysozyme and alpha-lactalbumin: structure, function, and interrelationships". Advances in Protein Chemistry. 41: 173–315. doi:10.1016/s0065-3233(08)60198-9. ISBN 9780120342419. PMID 2069076.
- Warshel A (November 1978). "Energetics of enzyme catalysis". Proceedings of the National Academy of Sciences of the United States of America. 75 (11): 5250–4. Bibcode:1978PNAS...75.5250W. doi:10.1073/pnas.75.11.5250. PMC 392938. PMID 281676.
- Weber JP, Fink AL (October 1980). "Temperature-dependent change in the rate-limiting step of beta-glucosidase catalysis". The Journal of Biological Chemistry. 255 (19): 9030–2. PMID 6773958.
- http://proteopedia.org/wiki/index.php/Lysozyme#Covalent_intermediate_and_product_complex
- Dahlquist FW, Rand-Meir T, Raftery MA (October 1969). "Application of secondary alpha-deuterium kinetic isotope effects to studies of enzyme catalysis. Glycoside hydrolysis by lysozyme and beta-glucosidase". Biochemistry. 8 (10): 4214–21. doi:10.1021/bi00838a045. PMID 5388150.
- Vocadlo DJ, Davies GJ, Laine R, Withers SG (August 2001). "Catalysis by hen egg-white lysozyme proceeds via a covalent intermediate" (PDF). Nature. 412 (6849): 835–8. Bibcode:2001Natur.412..835V. doi:10.1038/35090602. PMID 11518970.
- Grisham CM, Garrett RH (2007). "Chapter 14: Mechanism of enzyme action". Biochemistry. Australia: Thomson Brooks/Cole. pp. 467–9. ISBN 978-0-495-11912-8.
- Koshland DE (November 1953). "Stereochemistry and the Mechanism of Enzymatic Reactions". Biological Reviews. 28 (4): 416–436. doi:10.1111/j.1469-185X.1953.tb01386.x.
- Bowman AL, Grant IM, Mulholland AJ (October 2008). "QM/MM simulations predict a covalent intermediate in the hen egg white lysozyme reaction with its natural substrate". Chemical Communications (37): 4425–7. doi:10.1039/b810099c. PMID 18802578.
- Swan ID (March 1972). "The inhibition of hen egg-white lysozyme by imidazole and indole derivatives". Journal of Molecular Biology. 65 (1): 59–62. doi:10.1016/0022-2836(72)90491-3. PMID 5063023.
- Ohno N, Morrison DC (March 1989). "Lipopolysaccharide interaction with lysozyme. Binding of lipopolysaccharide to lysozyme and inhibition of lysozyme enzymatic activity". The Journal of Biological Chemistry. 264 (8): 4434–41. PMID 2647736.
- Ibrahim HR, Matsuzaki T, Aoki T (2001). "Genetic evidence that antibacterial activity of lysozyme is independent of its catalytic function". FEBS Letters. 506 (1): 27–32. doi:10.1016/S0014-5793(01)02872-1. PMID 11591365.
- Zhang R, Wu L, Eckert T, Burg-Roderfeld M, Rojas-Macias MA, Lütteke T (2017). "Lysozyme's lectin-like characteristics facilitates its immune defense function". Quarterly Reviews of Biophysics. 50: e9. doi:10.1017/S0033583517000075. PMID 29233221.
- Grivel JC, Smith-Gill SJ (1996). Lysozyme: Antigenic structure as defined by antibody and T cell responses. CRC Press. pp. 91–144. ISBN 978-0-8493-9225-2.
- Choi Y, Moody IS, Sims PC, Hunt SR, Corso BL, Perez I, Weiss GA, Collins PG (January 2012). "Single-molecule lysozyme dynamics monitored by an electronic circuit". Science. 335 (6066): 319–24. Bibcode:2012Sci...335..319C. doi:10.1126/science.1214824. PMC 3914775. PMID 22267809.
- Revenis ME, Kaliner MA (August 1992). "Lactoferrin and lysozyme deficiency in airway secretions: association with the development of bronchopulmonary dysplasia". The Journal of Pediatrics. 121 (2): 262–70. doi:10.1016/S0022-3476(05)81201-6. PMID 1640295.
- Cooper CA, Garas Klobas LC, Maga EA, Murray JD (2013). "Consuming transgenic goats' milk containing the antimicrobial protein lysozyme helps resolve diarrhea in young pigs". PLOS ONE. 8 (3): e58409. Bibcode:2013PLoSO...858409C. doi:10.1371/journal.pone.0058409. PMC 3596375. PMID 23516474.
- Molteni M (30 June 2016). "Spilled Milk". Case Studies: News Features. Undark: Truth, Beauty, Science. Retrieved 12 January 2017.
- Nester EW, Anderson DG, Roberts CE, Nester MT (2007). Microbiology: A Human Perspective (5th ed.). Boston, Mass.: McGraw-Hill Higher Education. ISBN 978-0-07-110706-8.
- Chandra RK (September 1978). "Immunological aspects of human milk". Nutrition Reviews. 36 (9): 265–72. doi:10.1111/j.1753-4887.1978.tb07393.x. PMID 362248.
- Tomita H, Sato S, Matsuda R, Sugiura Y, Kawaguchi H, Niimi T, Yoshida S, Morishita M (1999). "Serum lysozyme levels and clinical features of sarcoidosis". Lung. 177 (3): 161–7. doi:10.1007/pl00007637. PMID 10192763.
- Kenner GW (June 1977). "The Bakerian lecture. Towards synthesis of proteins". Proceedings of the Royal Society of London. Series B, Biological Sciences. 197 (1128): 237–53. Bibcode:1977RSPSB.197..237K. doi:10.1098/rspb.1977.0068. PMID 19745.
- Durek T, Torbeev VY, Kent SB (March 2007). "Convergent chemical synthesis and high-resolution x-ray structure of human lysozyme". Proceedings of the National Academy of Sciences of the United States of America. 104 (12): 4846–51. Bibcode:2007PNAS..104.4846D. doi:10.1073/pnas.0610630104. PMC 1829227. PMID 17360367.
- Wei H, Wang Z, Zhang J, House S, Gao YG, Yang L, Robinson H, Tan LH, Xing H, Hou C, Robertson IM, Zuo JM, Lu Y (February 2011). "Time-dependent, protein-directed growth of gold nanoparticles within a single crystal of lysozyme". Nature Nanotechnology. 6 (2): 93–7. Bibcode:2011NatNa...6...93W. doi:10.1038/nnano.2010.280. PMID 21278750.
- Sanghamitra NJ, Ueno T (May 2013). "Expanding coordination chemistry from protein to protein assembly". Chemical Communications. 49 (39): 4114–26. doi:10.1039/C2CC36935D. PMID 23211931.
- Ueno T (July 2013). "Porous protein crystals as reaction vessels". Chemistry. 19 (28): 9096–102. doi:10.1002/chem.201300250. PMID 23813903.
- Repaske R (October 1956). "Lysis of gram-negative bacteria by lysozyme". Biochimica et Biophysica Acta. 22 (1): 189–91. doi:10.1016/0006-3002(56)90240-2. PMID 13373865.
- Gunton J, Shiryayev A, Pagan DL (2007). Protein Condensation : Kinetic Pathways to Crystallization and Disease. Cambridge: Cambridge University Press. pp. 156–158. ISBN 9780511535321.
- Ninfa A, Ballou D, Benore M (2010). Fundamental Laboratory Approaches for Biochemistry and Biotechnology. John Wiley. ISBN 978-0470087664.
- Salton MR (June 1957). "The properties of lysozyme and its action on microorganisms". Bacteriological Reviews. 21 (2): 82–100. doi:10.1128/MMBR.21.2.82-100.1957. PMC 180888. PMID 13436356.
- Laschtschenko P (1909). "Über die keimtötende und entwicklungshemmende Wirkung Hühnereiweiß" [On the germ-killing and growth-inhibiting effect chicken egg albumin]. Z. Hyg. InfektKrankh. (in German). 64: 419–427. doi:10.1007/BF02216170.
- Duckett, S. (1999). "Duckett S. Ernest Duchesne and the concept of fungal antibiotic therapy. The Lancet Department Of Medical History 354(9195):2068-2071, December 11, 1999". Lancet. 354 (9195): 2068–71. doi:10.1016/S0140-6736(99)03162-1. PMID 10636385.
- Fleming A (May 1922). "On a remarkable bacteriolytic element found in tissues and secretions". Proceedings of the Royal Society B. 93 (653): 306–317. Bibcode:1922RSPSB..93..306F. doi:10.1098/rspb.1922.0023. JSTOR 80959.
- Advances in Protein Chemistry. Academic Press. 13 June 1991. pp. 176–. ISBN 978-0-08-058214-6.
- Blake CC, Koenig DF, Mair GA, North AC, Phillips DC, Sarma VR (May 1965). "Structure of hen egg-white lysozyme. A three-dimensional Fourier synthesis at 2 Angstrom resolution". Nature. 206 (4986): 757–61. doi:10.1038/206757a0. PMID 5891407.
- Johnson LN, Phillips DC (May 1965). "Structure of some crystalline lysozyme-inhibitor complexes determined by X-ray analysis at 6 Angstrom resolution". Nature. 206 (4986): 761–3. doi:10.1038/206761a0. PMID 5840126.
- Johnson LN (November 1998). "The early history of lysozyme". Nature Structural Biology. 5 (11): 942–4. doi:10.1038/2917. PMID 9808036.
- Canfield RE (August 1963). "The Amino Acid Sequence of Egg White Lysozyme". The Journal of Biological Chemistry. 238 (8): 2698–707. PMID 14063294.
- Vernon CA (April 1967). "The mechanisms of hydrolysis of glycosides and their relevance to enzyme-catalysed reactions". Proceedings of the Royal Society of London. Series B, Biological Sciences. 167 (1009): 389–401. Bibcode:1967RSPSB.167..389V. doi:10.1098/rspb.1967.0036. JSTOR 75680. PMID 4382802.
- Rupley JA (April 1967). "The binding and cleavage by lysozyme of N-acetylglucosamine oligosaccharides". Proceedings of the Royal Society of London. Series B, Biological Sciences. 167 (1009): 416–28. Bibcode:1967RSPSB.167..416R. doi:10.1098/rspb.1967.0038. JSTOR 75682. PMID 4382804.
- Sharon N (April 1967). "The chemical structure of lysozyme substrates and their cleavage by the enzyme". Proceedings of the Royal Society of London. Series B, Biological Sciences. 167 (1009): 402–15. Bibcode:1967RSPSB.167..402S. doi:10.1098/rspb.1967.0037. JSTOR 75681. PMID 4382803.
External links
- Muramidase at the US National Library of Medicine Medical Subject Headings (MeSH)
- Proteopedia.org HEW Lysozyme
- PDBe-KB provides an overview of all the structure information available in the PDB for Human Lysozyme C.
- PDBe-KB provides an overview of all the structure information available in the PDB for Hen egg white Lysozyme C.