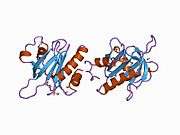
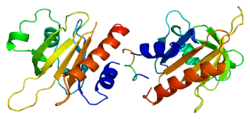Profilin 1
Profilin-1 is a protein that in humans is encoded by the PFN1 gene.[5][6]
Function
The protein encoded by this gene is a ubiquitous actin monomer-binding protein belonging to the profilin family. It is thought to regulate actin polymerization in response to extracellular signals. Deletion of this gene is associated with Miller-Dieker syndrome.[7] Mutations in this gene may be a rare cause of amyotrophic lateral sclerosis, also called Lou Gehrig's disease.[8][9][10][11][12][13][14][15][16][17][18][19][20]
Interactions
Profilin 1 has been shown to interact with:
gollark: I don't think you can say that "Any image which does not contain red does not contain red." is wrong, just unhelpful.
gollark: Oh, by red I mean not unred.
gollark: Any image which does not contain red does not contain red.
gollark: Sure, that might be "obviously a tautology" and "an unhelpful thing to say", but too bad.
gollark: Purely nonred colors.
References
- GRCh38: Ensembl release 89: ENSG00000108518 - Ensembl, May 2017
- GRCm38: Ensembl release 89: ENSMUSG00000018293 - Ensembl, May 2017
- "Human PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- "Mouse PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- Kwiatkowski DJ, Bruns GA (May 1988). "Human profilin. Molecular cloning, sequence comparison, and chromosomal analysis". J Biol Chem. 263 (12): 5910–5. PMID 3356709.
- Kwiatkowski DJ, Aklog L, Ledbetter DH, Morton CC (April 1990). "Identification of the functional profilin gene, its localization to chromosome subband 17p13.3, and demonstration of its deletion in some patients with Miller-Dieker syndrome". Am J Hum Genet. 46 (3): 559–67. PMC 1683621. PMID 1968707.
- "Entrez Gene: PFN1 profilin 1".
- Wu CH, Fallini C, Ticozzi N, Keagle PJ, Sapp PC, Piotrowska K, et al. (August 2012). "Mutations in the profilin 1 gene cause familial amyotrophic lateral sclerosis". Nature. 488 (7412): 499–503. doi:10.1038/nature11280. PMC 3575525. PMID 22801503.
- Daoud H, Dobrzeniecka S, Camu W, Meininger V, Dupré N, Dion PA, Rouleau GA (April 2013). "Mutation analysis of PFN1 in familial amyotrophic lateral sclerosis patients". Neurobiology of Aging. 34 (4): 1311.e1-2. doi:10.1016/j.neurobiolaging.2012.09.001. PMID 23062600.
- Tiloca C, Ticozzi N, Pensato V, Corrado L, Del Bo R, Bertolin C, et al. (May 2013). "Screening of the PFN1 gene in sporadic amyotrophic lateral sclerosis and in frontotemporal dementia". Neurobiology of Aging. 34 (5): 1517.e9-10. doi:10.1016/j.neurobiolaging.2012.09.016. PMC 3548975. PMID 23063648.
- Ingre C, Landers JE, Rizik N, Volk AE, Akimoto C, Birve A, et al. (June 2013). "A novel phosphorylation site mutation in profilin 1 revealed in a large screen of US, Nordic, and German amyotrophic lateral sclerosis/frontotemporal dementia cohorts". Neurobiology of Aging. 34 (6): 1708.e1-6. doi:10.1016/j.neurobiolaging.2012.10.009. PMC 6591725. PMID 23141414.
- Lattante S, Le Ber I, Camuzat A, Brice A, Kabashi E (June 2013). "Mutations in the PFN1 gene are not a common cause in patients with amyotrophic lateral sclerosis and frontotemporal lobar degeneration in France". Neurobiology of Aging. 34 (6): 1709.e1-2. doi:10.1016/j.neurobiolaging.2012.10.026. PMID 23182804.
- Dillen L, Van Langenhove T, Engelborghs S, Vandenbulcke M, Sarafov S, Tournev I, et al. (June 2013). "Explorative genetic study of UBQLN2 and PFN1 in an extended Flanders-Belgian cohort of frontotemporal lobar degeneration patients". Neurobiology of Aging. 34 (6): 1711.e1-5. doi:10.1016/j.neurobiolaging.2012.12.007. PMID 23312802.
- Zou ZY, Sun Q, Liu MS, Li XG, Cui LY (June 2013). "Mutations in the profilin 1 gene are not common in amyotrophic lateral sclerosis of Chinese origin". Neurobiology of Aging. 34 (6): 1713.e5-6. doi:10.1016/j.neurobiolaging.2012.12.024. PMID 23357624.
- Chen Y, Zheng ZZ, Huang R, Chen K, Song W, Zhao B, et al. (July 2013). "PFN1 mutations are rare in Han Chinese populations with amyotrophic lateral sclerosis". Neurobiology of Aging. 34 (7): 1922.e1-5. doi:10.1016/j.neurobiolaging.2013.01.013. PMID 23428184.
- van Blitterswijk M, Baker MC, Bieniek KF, Knopman DS, Josephs KA, Boeve B, et al. (September 2013). "Profilin-1 mutations are rare in patients with amyotrophic lateral sclerosis and frontotemporal dementia". Amyotrophic Lateral Sclerosis & Frontotemporal Degeneration. 14 (5–6): 463–9. doi:10.3109/21678421.2013.787630. PMC 3923463. PMID 23634771.
- Yang S, Fifita JA, Williams KL, Warraich ST, Pamphlett R, Nicholson GA, et al. (September 2013). "Mutation analysis and immunopathological studies of PFN1 in familial and sporadic amyotrophic lateral sclerosis". Neurobiology of Aging. 34 (9): 2235.e7-10. doi:10.1016/j.neurobiolaging.2013.04.003. PMID 23635659.
- Fratta P, Charnock J, Collins T, Devoy A, Howard R, Malaspina A, et al. (May 2014). "Profilin1 E117G is a moderate risk factor for amyotrophic lateral sclerosis". Journal of Neurology, Neurosurgery, and Psychiatry. 85 (5): 506–8. doi:10.1136/jnnp-2013-306761. PMC 3995330. PMID 24309268.
- Syriani E, Salvans C, Salvadó M, Morales M, Lorenzo L, Cazorla S, Gamez J (December 2014). "PFN1 mutations are also rare in the Catalan population with amyotrophic lateral sclerosis". Journal of Neurology. 261 (12): 2387–92. doi:10.1007/s00415-014-7501-x. PMID 25249294.
- Smith BN, Vance C, Scotter EL, Troakes C, Wong CH, Topp S, et al. (March 2015). "Novel mutations support a role for Profilin 1 in the pathogenesis of ALS". Neurobiology of Aging. 36 (3): 1602.e17-27. doi:10.1016/j.neurobiolaging.2014.10.032. PMC 4357530. PMID 25499087.
- Yayoshi-Yamamoto S, Taniuchi I, Watanabe T (September 2000). "FRL, a novel formin-related protein, binds to Rac and regulates cell motility and survival of macrophages". Mol. Cell. Biol. 20 (18): 6872–81. doi:10.1128/mcb.20.18.6872-6881.2000. PMC 86228. PMID 10958683.
- Boettner B, Govek EE, Cross J, Van Aelst L (August 2000). "The junctional multidomain protein AF-6 is a binding partner of the Rap1A GTPase and associates with the actin cytoskeletal regulator profilin". Proc. Natl. Acad. Sci. U.S.A. 97 (16): 9064–9. doi:10.1073/pnas.97.16.9064. PMC 16822. PMID 10922060.
- Harbeck B, Hüttelmaier S, Schluter K, Jockusch BM, Illenberger S (October 2000). "Phosphorylation of the vasodilator-stimulated phosphoprotein regulates its interaction with actin". J. Biol. Chem. 275 (40): 30817–25. doi:10.1074/jbc.M005066200. PMID 10882740.
- Miki H, Suetsugu S, Takenawa T (December 1998). "WAVE, a novel WASP-family protein involved in actin reorganization induced by Rac". EMBO J. 17 (23): 6932–41. doi:10.1093/emboj/17.23.6932. PMC 1171041. PMID 9843499.
- Mimuro H, Suzuki T, Suetsugu S, Miki H, Takenawa T, Sasakawa C (September 2000). "Profilin is required for sustaining efficient intra- and intercellular spreading of Shigella flexneri". J. Biol. Chem. 275 (37): 28893–901. doi:10.1074/jbc.M003882200. PMID 10867004.
- Suetsugu S, Miki H, Takenawa T (November 1998). "The essential role of profilin in the assembly of actin for microspike formation". EMBO J. 17 (22): 6516–26. doi:10.1093/emboj/17.22.6516. PMC 1170999. PMID 9822597.
Further reading
- Qualmann B, Kessels MM (2003). "Endocytosis and the cytoskeleton". Int. Rev. Cytol. International Review of Cytology. 220: 93–144. doi:10.1016/S0074-7696(02)20004-2. ISBN 978-0-12-364624-8. PMID 12224553.
- Ampe C, Markey F, Lindberg U, Vandekerckhove J (1988). "The primary structure of human platelet profilin: reinvestigation of the calf spleen profilin sequence". FEBS Lett. 228 (1): 17–21. doi:10.1016/0014-5793(88)80575-1. PMID 3342873.
- Gieselmann R, Kwiatkowski DJ, Janmey PA, Witke W (1995). "Distinct biochemical characteristics of the two human profilin isoforms". Eur. J. Biochem. 229 (3): 621–8. doi:10.1111/j.1432-1033.1995.tb20506.x. PMID 7758455.
- Kato S, Sekine S, Oh SW, Kim NS, Umezawa Y, Abe N, Yokoyama-Kobayashi M, Aoki T (1995). "Construction of a human full-length cDNA bank". Gene. 150 (2): 243–50. doi:10.1016/0378-1119(94)90433-2. PMID 7821789.
- Metzler WJ, Constantine KL, Friedrichs MS, Bell AJ, Ernst EG, Lavoie TB, Mueller L (1994). "Characterization of the three-dimensional solution structure of human profilin: 1H, 13C, and 15N NMR assignments and global folding pattern". Biochemistry. 32 (50): 13818–29. doi:10.1021/bi00213a010. PMID 8268157.
- Schutt CE, Myslik JC, Rozycki MD, Goonesekere NC, Lindberg U (1993). "The structure of crystalline profilin-beta-actin". Nature. 365 (6449): 810–6. doi:10.1038/365810a0. PMID 8413665.
- Mahoney NM, Janmey PA, Almo SC (1997). "Structure of the profilin-poly-L-proline complex involved in morphogenesis and cytoskeletal regulation". Nat. Struct. Biol. 4 (11): 953–60. doi:10.1038/nsb1197-953. PMID 9360613.
- Mammoto A, Sasaki T, Asakura T, Hotta I, Imamura H, Takahashi K, Matsuura Y, Shirao T, Takai Y (1998). "Interactions of drebrin and gephyrin with profilin". Biochem. Biophys. Res. Commun. 243 (1): 86–9. doi:10.1006/bbrc.1997.8068. PMID 9473484.
- Bhargavi V, Chari VB, Singh SS (1998). "Phosphatidylinositol 3-kinase binds to profilin through the p85 alpha subunit and regulates cytoskeletal assembly". Biochem. Mol. Biol. Int. 46 (2): 241–8. doi:10.1080/15216549800203752. PMID 9801792.
- Suetsugu S, Miki H, Takenawa T (1999). "The essential role of profilin in the assembly of actin for microspike formation". EMBO J. 17 (22): 6516–26. doi:10.1093/emboj/17.22.6516. PMC 1170999. PMID 9822597.
- Miki H, Suetsugu S, Takenawa T (1999). "WAVE, a novel WASP-family protein involved in actin reorganization induced by Rac". EMBO J. 17 (23): 6932–41. doi:10.1093/emboj/17.23.6932. PMC 1171041. PMID 9843499.
- Mahoney NM, Rozwarski DA, Fedorov E, Fedorov AA, Almo SC (1999). "Profilin binds proline-rich ligands in two distinct amide backbone orientations". Nat. Struct. Biol. 6 (7): 666–71. doi:10.1038/10722. PMID 10404225.
- Nunoi H, Yamazaki T, Tsuchiya H, Kato S, Malech HL, Matsuda I, Kanegasaki S (1999). "A heterozygous mutation of β-actin associated with neutrophil dysfunction and recurrent infection". Proc. Natl. Acad. Sci. U.S.A. 96 (15): 8693–8. doi:10.1073/pnas.96.15.8693. PMC 17578. PMID 10411937.
- Murphy GA, Solski PA, Jillian SA, Pérez de la Ossa P, D'Eustachio P, Der CJ, Rush MG (1999). "Cellular functions of TC10, a Rho family GTPase: regulation of morphology, signal transduction and cell growth". Oncogene. 18 (26): 3831–45. doi:10.1038/sj.onc.1202758. PMID 10445846.
- Harbeck B, Hüttelmaier S, Schluter K, Jockusch BM, Illenberger S (2000). "Phosphorylation of the vasodilator-stimulated phosphoprotein regulates its interaction with actin". J. Biol. Chem. 275 (40): 30817–25. doi:10.1074/jbc.M005066200. PMID 10882740.
- Boettner B, Govek EE, Cross J, Van Aelst L (2000). "The junctional multidomain protein AF-6 is a binding partner of the Rap1A GTPase and associates with the actin cytoskeletal regulator profilin". Proc. Natl. Acad. Sci. U.S.A. 97 (16): 9064–9. doi:10.1073/pnas.97.16.9064. PMC 16822. PMID 10922060.
- Yayoshi-Yamamoto S, Taniuchi I, Watanabe T (2000). "FRL, a Novel Formin-Related Protein, Binds to Rac and Regulates Cell Motility and Survival of Macrophages". Mol. Cell. Biol. 20 (18): 6872–81. doi:10.1128/MCB.20.18.6872-6881.2000. PMC 86228. PMID 10958683.
- Mellon MB, Frank BT, Fang KC (2002). "Mast cell alpha-chymase reduces IgE recognition of birch pollen profilin by cleaving antibody-binding epitopes". J. Immunol. 168 (1): 290–7. doi:10.4049/jimmunol.168.1.290. PMID 11751973.
This article is issued from Wikipedia. The text is licensed under Creative Commons - Attribution - Sharealike. Additional terms may apply for the media files.