Epithelial cell adhesion molecule
Epithelial cell adhesion molecule (EpCAM) is a transmembrane glycoprotein mediating Ca2+-independent homotypic cell–cell adhesion in epithelia.[5] EpCAM is also involved in cell signaling,[6] migration,[7] proliferation, and differentiation.[8] Additionally, EpCAM has oncogenic potential via its capacity to upregulate c-myc, e-fabp, and cyclins A & E.[9] Since EpCAM is expressed exclusively in epithelia and epithelial-derived neoplasms, EpCAM can be used as diagnostic marker for various cancers. It appears to play a role in tumorigenesis and metastasis of carcinomas, so it can also act as a potential prognostic marker and as a potential target for immunotherapeutic strategies.[10]
Expression pattern
First discovered in 1979, EpCAM was initially described as a dominant surface antigen on human colon carcinoma.[11] Because of its prevalence on many carcinomas, it has been "discovered" many different times.[12] EpCAM therefore has many aliases the most notable of which include TACSTD1 (tumor-associated calcium signal transducer 1), CD326 (cluster of differentiation 326), and the 17-1A antigen.[13]
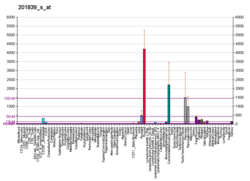
EpCAM expression is not limited to human colon carcinomas; in fact, EpCAM is expressed in a variety of human epithelial tissues, carcinomas, and progenitor and stem cells. However, EpCAM is not found in non-epithelial cells or cancers of non-epithelial origin. EpCAM is expressed on the basolateral membrane of all simple (especially glandular), pseudo-stratified, and transitional epithelia. In contrast, normal squamous stratified epithelia are negative for EpCAM. The level of expression may differ significantly between the individual tissue types. In the gastrointestinal tract, the gastric epithelium expresses very low levels of EpCAM. Expression levels are substantially higher in small intestine, and in colon EpCAM is probably expressed at the highest levels among all epithelial cell types.[13]
EpCAM is frequently upregulated in carcinomas but is not expressed in cancers of non-epithelial origin. In cancer cells, EpCAM is expressed in a dispersed pattern across the cell membrane.[14] However, EpCAM expression in carcinomas is often heterogeneous; some cells in a tumor have more EpCAM than other cells in the same tumor.
Squamous carcinomas often express EpCAM whereas normal squamous cells do not express EpCAM. EpCAM expression differs between different types of renal cell carcinomas, and EpCAM expression increases during development of androgen resistance in prostate cancer.[15] All of this points towards the utility of EpCAM as a diagnostic tool for various cancers.
Structure
Although it is identified as a cell adhesion molecule, EpCAM does not structurally resemble any of the four major families of cell adhesion molecules, namely cadherins, integrins, selectins, and members of the immunoglobulin super-family.[13]
EpCAM is a glycosylated, 30- to 40-kDa type I membrane protein. The sequence of the EpCAM molecule predicts the presence of three potential N-linked glycosylation sites. It is composed of 314 amino acids. EpCAM consists of an extracellular domain (242 amino acids) with epidermal growth factor (EGF)- and thyroglobulin repeat-like domains, a single transmembrane domain (23 amino acids), and a short intracellular domain (26 amino acids).[10] The extracellular domain is sometimes referred to as EpEX, and the intracellular domain is sometimes referred to as EpICD.[14]
Function
The exact function of EpCAM is currently being elucidated, but EpCAM appears to play many different roles.
Cell adhesion
EpCAM was first found to play a role in homotypic cell adhesion.[5] This means that EpCAM on the surface of one cell binds to the EpCAM on a neighboring cell thereby holding the cells together. The adhesions mediated by EpCAM are relatively weak, as compared to some other adhesion molecules, such as classic cadherins.
EpICD is required for EpCAM to mediate intercellular adhesion; EpCAM mediates intercellular adhesion and associates with the actin cytoskeleton via EpICD.[16]
EpCAM has a negative impact on cadherin-mediated adhesions. Overexpression of EpCAM does not alter the expression level of cadherin but rather decreases the association of the cadherin/ catenin complex in the cytoskeleton. As EpCAM expression increases, the total amount of α-catenin decreases, whereas cellular β-catenin levels remain constant.[17] Active proliferation in a number of epithelial tissues is associated with increased or de novo EpCAM expression. This is especially evident in tissues that normally reveal no or low levels of EpCAM expression, such as squamous epithelium. The level of EpCAM expression correlates with the proliferative activity of intestinal cells, and inversely correlates with their differentiation.[8]
Role in cancer
EpCAM can be cleaved which lends the molecule oncogenic potential. Upon cleavage, the extracellular domain (EpEX) is released into the area surrounding the cell, and the intracellular domain (EpICD) is released into the cytoplasm of the cell. EpICD forms a complex with the proteins FHL2, β-catenin, and Lef inside the nucleus. This complex then binds to DNA and promotes the transcription of various genes. Targets of upregulation include c-myc, e-fabp, and cyclins A & E.[6] This has the effect of promoting tumor growth. Additionally, EpEX that has been cleaved can stimulate the cleavage of additional EpCAM molecules resulting in a positive feedback loop.[14] The amount of β-catenin in the nucleus can modulate the expression level of EpCAM.[18]
EpCAM may also play a role in epithelial mesenchymal transition (EMT) in tumors, although its exact effects are poorly understood. Its ability to suppress E-cadherin suggests that EpCAM would promote EMT and tumor metastasis, but its homotypic cell adhesion properties can counteract its ability to suppress E-cadherin.[19] Results from different studies are often conflicting. In one study, for example, silencing of EpCAM with short interfering RNA (siRNA) led to a reduction of proliferation, migration, and invasion of breast cancer cells in vitro[7] supporting the role of EpCAM in promoting EMT. In another study, cells undergoing EMT were found to downregulate EpCAM.[20] In one study, epithelial tumors were often strongly positive for EpCAM, but mesenchymal tumors showed only occasional and weak positivity.[15] It has been suggested that EpCAM expression is downregulated during EMT but then upregulated once the metastasis reaches its future tumor site.[21]
Clinical significance
Target for immunotherapy
It has been speculated that since EpCAM in normal epithelia is expressed mostly on the basolateral membrane, it would be much less accessible to antibodies than EpCAM in cancer tissue, where it is homogeneously distributed on the cancer cell surface. In addition to being overexpressed in many carcinomas, EpCAM is expressed in cancer stem cells, making EpCAM an attractive target for immunotherapy. However, the heterogeneous expression of EpCAM in carcinomas and the fact that EpCAM is not tumor-specific (i.e., it is found in normal epithelium) raise concerns that immunotherapy directed towards EpCAM could have severe side effects.[13] As the role of EpCAM in cancer cell signaling is better understood, EpCAM signaling rather than EpCAM itself may be a target for therapeutic intervention.[14]
Edrecolomab, catumaxomab and other monoclonal antibodies are designed to bind to it.[10][22] also nofetumomab.
Histopathology
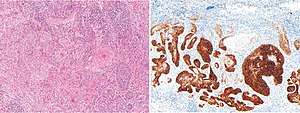
EpCAM is often overexpressed in certain carcinomas, including in breast cancer, colon cancer and basal cell carcinoma of the skin.[24] The diagnosis of such conditions can therefore be assisted by immunohistochemistry using BerEp4, which is an antibody to EpCAM.[24]
Genetic disorders
A problem in EpCAM can indirectly cause Lynch syndrome,[25] a genetic disorder that leads to increased risk of cancer. Deletion of a portion of the 3' end of the EpCAM gene causes epigenetic inactivation of the MSH2 gene by hypermethylating the promoter region of the MSH2 gene.
Mutations in EpCAM have also been associated with congenital tufting enteropathy[26] which causes intractable diarrhea in newborn children.
References
- GRCh38: Ensembl release 89: ENSG00000119888 - Ensembl, May 2017
- GRCm38: Ensembl release 89: ENSMUSG00000045394 - Ensembl, May 2017
- "Human PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- "Mouse PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- Litvinov, Sergey; et al. (1994). "Ep-CAM: a human epithelial antigen is a homophilic cell–cell adhesion molecule". The Journal of Cell Biology. 125 (2): 437–46. doi:10.1083/jcb.125.2.437. PMC 2120036. PMID 8163559.
- Maetzel, Dorothea; et al. (2009). "Nuclear signalling by tumour-associated antigen EpCAM". Nature Cell Biology. 11 (2): 162–71. doi:10.1038/ncb1824. PMID 19136966.
- Osta, WA; et al. (2004). "EpCAM is overexpressed in breast cancer and is a potential target for breast cancer gene therapy". Cancer Res. 64 (16): 5818–24. doi:10.1158/0008-5472.CAN-04-0754. PMID 15313925.
- Litvinov, Sergey; et al. (1996). "Expression of Ep-CAM in cervical squamous epithelia correlates with an increased proliferation and the disappearance of markers for terminal differentiation". Am J Pathol. 148 (3): 865–75. PMC 1861708. PMID 8774141.
- Munz, M; et al. (2004). "The carcinoma-associated antigen EpCAM upregulates c-myc and induces cell proliferation". Oncogene. 23 (34): 5748–58. doi:10.1038/sj.onc.1207610. PMID 15195135.
- Armstrong, Andrew; Stephen Eck (2003). "EpCAM: a new therapeutic target for an old cancer antigen". Cancer Biology & Therapy. 2 (4): 320–6. doi:10.4161/cbt.2.4.451. PMID 14508099.
- Herlyn, D; et al. (1979). "Monoclonal antibodies in cell-mediated cytotoxicity against human melanoma and colorectal carcinoma". Eur J Immunol. 9 (8): 657–9. doi:10.1002/eji.1830090817. PMID 499332.
- Baeuerle, P. A.; Gires O. (2007). "EpCAM (CD326) finding its role in cancer". Br J Cancer. 96 (3): 417–23. doi:10.1038/sj.bjc.6603494. PMC 2360029. PMID 17211480.
- Balzar, M; Winter MJ; de Boer CJ; Litvinov SV (October 1999). "The biology of the 17-1A antigen (Ep-CAM)". J. Mol. Med. 77 (10): 699–712. doi:10.1007/s001099900038. PMID 10606205.
- Munz, M; P. A. Baeuerle; O. Gires (2009). "The emerging role of EpCAM in cancer and stem cell signaling". Cancer Research. 69 (14): 5627–9. doi:10.1158/0008-5472.CAN-09-0654. PMID 19584271.
- Went, PT; et al. (2004). "Frequent EpCam protein expression in human carcinomas". Hum Pathol. 35 (1): 122–8. doi:10.1016/j.humpath.2003.08.026. PMID 14745734.
- Balzar, M; Bakker HA; Briaire-de-Bruijn IH; Fleuren GJ; Warnaar SO; Litvinov SV (1998). "Cytoplasmic tail regulates the intercellular adhesion function of the epithelial cell adhesion molecule". Mol Cell Biol. 18 (8): 4833–43. doi:10.1128/MCB.18.8.4833. PMC 109068. PMID 9671492.
- Litvinov, S.; et al. (1997). "Epithelial cell adhesion molecule (Ep-CAM) modulates cell–cell interactions mediated by classic cadherins". J Cell Biol. 139 (5): 1337–48. doi:10.1083/jcb.139.5.1337. PMC 2140211. PMID 9382878.
- Yamashita, T; Budhu A; Forgues M; Wang XW (2007). "Activation of hepatic stem cell marker EpCAM by Wnt–β-catenin signaling in hepatocellular carcinoma". Cancer Res. 67 (22): 10831–9. doi:10.1158/0008-5472.CAN-07-0908. PMID 18006828.
- Patriarca, C.; et al. (2012). "Epithelial cell adhesion molecule expression (CD326) in cancer: a short review". Cancer Treat Rev. 38 (1): 68–75. doi:10.1016/j.ctrv.2011.04.002. PMID 21576002.
- Santisteban, M.; et al. (2009). "Immune-induced epithelial to mesenchymal transition in vivo generates breast cancer stem cells". Cancer Res. 69 (7): 2887–95. doi:10.1158/0008-5472.CAN-08-3343. PMC 2664865. PMID 19276366.
- van der Gun, BT; et al. (2010). "EpCAM in carcinogenesis: the good, the bad or the ugly". Carcinogenesis. 31 (11): 1913–21. doi:10.1093/carcin/bgq187. PMID 20837599.
- Punt CJ, Nagy A, Douillard JY, Figer A, Skovsgaard T, Monson J, Barone C, Fountzilas G, Riess H, Moylan E, Jones D, Dethling J, Colman J, Coward L, MacGregor S (August 2002). "Edrecolomab alone or in combination with fluorouracil and folinic acid in the adjuvant treatment of stage III colon cancer: a randomised study". Lancet. 360 (9334): 671–7. doi:10.1016/S0140-6736(02)09836-7. PMID 12241873.
- Sunjaya, Anthony Paulo; Sunjaya, Angela Felicia; Tan, Sukmawati Tansil (2017). "The Use of BEREP4 Immunohistochemistry Staining for Detection of Basal Cell Carcinoma". Journal of Skin Cancer. 2017: 1–10. doi:10.1155/2017/2692604. ISSN 2090-2905.
- Dasgeb, Bahar; Mohammadi, Tarana M; Mehregan, Darius R (2013). "Use of Ber-EP4 and Epithelial Specific Antigen to Differentiate Clinical Simulators of Basal Cell Carcinoma". Biomarkers in Cancer. 5: BIC.S11856. doi:10.4137/BIC.S11856. ISSN 1179-299X.
- Tomita, N; Yamano T; Matsubara N; Tamura K (2013). "[A Novel Genetic Disorder of Lynch Syndrome-EPCAM Gene Deletion]". Gan to Kagaku Ryoho. 40 (2): 143–7. PMID 23411950.
- Sivagnanam, M; et al. (2008). "Identification of EpCAM as the gene for congenital tufting enteropathy". Gastroenterology. 135 (2): 429–37. doi:10.1053/j.gastro.2008.05.036. PMC 2574708. PMID 18572020.
External links
- FAQs on HNPCC from the National Institute of Health
- GeneReviews/NCBI/NIH/UW entry on Lynch syndrome
- TACSTD1+protein,+human at the US National Library of Medicine Medical Subject Headings (MeSH)
This article incorporates text from the United States National Library of Medicine, which is in the public domain.