Cellulase
Cellulase is any of several enzymes produced chiefly by fungi, bacteria, and protozoans that catalyze cellulolysis, the decomposition of cellulose and of some related polysaccharides. The name is also used for any naturally occurring mixture or complex of various such enzymes, that act serially or synergistically to decompose cellulosic material.
| Cellulase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
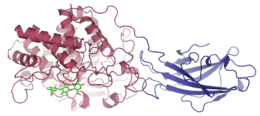 A cellulase enzyme produced by Thermomonospora fusca, with cellotriose bound in the shallow groove of the catalytic domain | |||||||||
| Identifiers | |||||||||
| EC number | 3.2.1.4 | ||||||||
| CAS number | 9012-54-8 | ||||||||
| Databases | |||||||||
| IntEnz | IntEnz view | ||||||||
| BRENDA | BRENDA entry | ||||||||
| ExPASy | NiceZyme view | ||||||||
| KEGG | KEGG entry | ||||||||
| MetaCyc | metabolic pathway | ||||||||
| PRIAM | profile | ||||||||
| PDB structures | RCSB PDB PDBe PDBsum | ||||||||
| Gene Ontology | AmiGO / QuickGO | ||||||||
| |||||||||
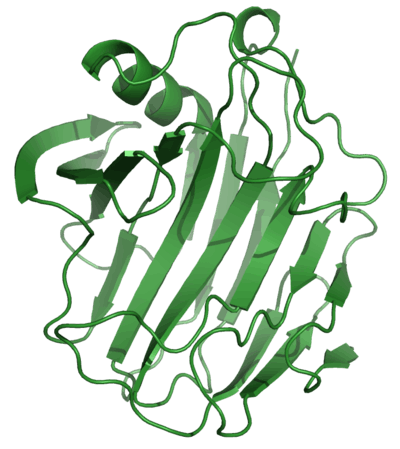
Cellulases break down the cellulose molecule into monosaccharides ("simple sugars") such as beta-glucose, or shorter polysaccharides and oligosaccharides. Cellulose breakdown is of considerable economic importance, because it makes a major constituent of plants available for consumption and use in chemical reactions. The specific reaction involved is the hydrolysis of the 1,4-beta-D-glycosidic linkages in cellulose, hemicellulose, lichenin, and cereal beta-D-glucans. Because cellulose molecules bind strongly to each other, cellulolysis is relatively difficult compared to the breakdown of other polysaccharides such as starch.[2]
Most mammals have only very limited ability to digest dietary fibres like cellulose by themselves. In many herbivorous animals such as ruminants like cattle and sheep and hindgut fermenters like horses, cellulases are produced by symbiotic bacteria. Endogenous cellulases are produced by a few types of metazoan animals, such as some termites, snails,[3][4][5] and earthworms.
Recently, cellulases have also been found in green microalgae (Chlamydomonas reinhardtii, Gonium pectorale and Volvox carteri) and their catalytic domains (CD) belonging to GH9 Family show highest sequence homology to metazoan endogenous cellulases. Algal cellulases are modular, consisting of putative novel cysteine-rich carbohydrate-binding modules (CBMs), proline/serine-(PS) rich linkers in addition to putative Ig-like and unknown domains in some members. Cellulase from Gonium pectorale consisted of two CDs separated by linkers and with a C-terminal CBM.[6]
Several different kinds of cellulases are known, which differ structurally and mechanistically. Synonyms, derivatives, and specific enzymes associated with the name "cellulase" include endo-1,4-beta-D-glucanase (beta-1,4-glucanase, beta-1,4-endoglucan hydrolase, endoglucanase D, 1,4-(1,3,1,4)-beta-D-glucan 4-glucanohydrolase), carboxymethyl cellulase (CMCase), avicelase, celludextrinase, cellulase A, cellulosin AP, alkali cellulase, cellulase A 3, 9.5 cellulase, and pancellase SS. Enzymes that cleave lignin have occasionally been called cellulases, but this old usage is deprecated; they are lignin-modifying enzymes.
Types and action
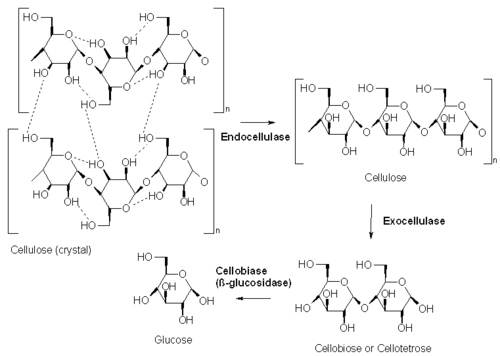
Five general types of cellulases based on the type of reaction catalyzed:
- Endocellulases (EC 3.2.1.4) randomly cleave internal bonds at amorphous sites that create new chain ends.
- Exocellulases or cellobiohydrolases (EC 3.2.1.91) cleave two to four units from the ends of the exposed chains produced by endocellulase, resulting in tetrasaccharides[7] or disaccharides, such as cellobiose. Exocellulases are further classified into type I, that work processively from the reducing end of the cellulose chain, and type II, that work processively from the nonreducing end.
- Cellobiases (EC 3.2.1.21) or beta-glucosidases hydrolyse the exocellulase product into individual monosaccharides.
- Oxidative cellulases depolymerize cellulose by radical reactions, as for instance cellobiose dehydrogenase (acceptor).
- Cellulose phosphorylases depolymerize cellulose using phosphates instead of water.
Avicelase has almost exclusively exo-cellulase activity, since avicel is a highly micro-crystalline substrate.
Within the above types there are also progressive (also known as processive) and nonprogressive types. Progressive cellulase will continue to interact with a single polysaccharide strand, nonprogressive cellulase will interact once then disengage and engage another polysaccharide strand.
Cellulase action is considered to be synergistic as all three classes of cellulase can yield much more sugar than the addition of all three separately. Aside from ruminants, most animals (including humans) do not produce cellulase in their bodies and can only partially break down cellulose through fermentation, limiting their ability to use energy in fibrous plant material.
Structure
Most fungal cellulases have a two-domain structure, with one catalytic domain and one cellulose binding domain, that are connected by a flexible linker. This structure is adapted for working on an insoluble substrate, and it allows the enzyme to diffuse two-dimensionally on a surface in a caterpillar-like fashion. However, there are also cellulases (mostly endoglucanases) that lack cellulose binding domains.
Both binding of substrates and catalysis depend on the three-dimensional structure of the enzyme which arises as a consequence of the level of protein folding. The amino acid sequence and arrangement of their residues that occur within the active site, the position where the substrate binds, may influence factors like binding affinity of ligands, stabilization of substrates within the active site and catalysis. The substrate structure is complementary to the precise active site structure of enzyme. Changes in the position of residues may result in distortion of one or more of these interactions.[8] Additional factors like temperature, pH and metal ions influence the non-covalent interactions between enzyme structure.[9] The Thermotoga maritima species make cellulases consisting of 2 beta-sheets (protein structures) surrounding a central catalytic region which is the active-site.[10] The enzyme is categorised as an endoglucanase, which internally cleaves β-1,4 -glycosydic bonds in cellulose chains facilitating further degradation of the polymer. Different species in the same family as T. Maritima make cellulases with different structures.[10] Cellulases produced by the species Coprinopsis Cinerea consists of seven protein strands in the shape of an enclosed tunnel called a beta/alpha barrel.[11] These enzymes hydrolyse the substrate carboxymethyl cellulose. Binding of the substrate in the active site induces a change in conformation which allows degradation of the molecule.
Cellulase complexes
In many bacteria, cellulases in-vivo are complex enzyme structures organized in supramolecular complexes, the cellulosomes. They can contain, but are not limited to, five different enzymatic subunits representing namely endocellulases, exocellulases, cellobiases, oxidative cellulases and cellulose phosphorylases wherein only endocellulases and cellobiases participate in the actual hydrolysis of the β(1→ 4) linkage. The number of sub-units making up cellulosomes can also determine the rate of enzyme activity.[12]
Multidomain cellulases are widespread among many taxonomic groups, however, cellulases from anaerobic bacteria, found in cellulosomes, have the most complex architecture consisting of different types of modules. For example, Clostridium cellulolyticum produces 13 GH9 modular cellulases containing a different number and arrangement of catalytic-domain (CD), carbohydrate-binding module (CBM), dockerin, linker and Ig-like domain.[13]
The cellulase complex from Trichoderma reesei, for example, comprises a component labeled C1 (57,000 daltons) that separates the chains of crystalline cellulose, an endoglucanase (about 52,000 daltons), an exoglucanase (about 61,000 dalton), and a beta-glucosidase (76,000 daltons).[14]
Numerous "signature" sequences known as dockerins and cohesins have been identified in the genomes of bacteria that produce cellulosomes. Depending on their amino acid sequence and tertiary structures, cellulases are divided into clans and families.[15]
Multimodular cellulases are more efficient than free enzyme (with only CD) due to synergism because of the close proximity between the enzyme and the cellulosic substrate. CBM are involved in binding of cellulose whereas glycosylated linkers provide flexibility to the CD for higher activity and protease protection, as well as increased binding to the cellulose surface.[6]
Mechanism of cellulolysis
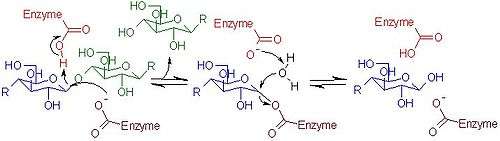
Uses
Cellulase is used for commercial food processing in coffee. It performs hydrolysis of cellulose during drying of beans. Furthermore, cellulases are widely used in textile industry and in laundry detergents. They have also been used in the pulp and paper industry for various purposes, and they are even used for pharmaceutical applications. Cellulase is used in the fermentation of biomass into biofuels, although this process is relatively experimental at present. Medically, Cellulase is used as a treatment for phytobezoars, a form of cellulose bezoar found in the human stomach, and it has exhibited efficacy in degrading polymicrobial bacterial biofilms by hydrolyzing the β(1-4) glycosidic linkages within the structural, matrix exopolysaccharides of the extracellular polymeric substance (EPS).[17][18]
Measurement
As the native substrate, cellulose, is a water-insoluble polymer, traditional reducing sugar assays using this substrate can not be employed for the measurement of cellulase activity. Analytical scientists have developed a number of alternative methods.
- DNSA Method Cellulase activity was determined by incubating 0.5 ml of supernatant with 0.5 ml of 1% carboxymethylcellulose (CMC) in 0.05M citrate buffer (pH 4.8) at 50°C for 30 minutes. The reaction was terminated by the addition of 3 ml dinitrosalicylic acid reagent. Absorbance was read at 540 nm.[19]
A viscometer can be used to measure the decrease in viscosity of a solution containing a water-soluble cellulose derivative such as carboxymethyl cellulose upon incubation with a cellulase sample.[20] The decrease in viscosity is directly proportional to the cellulase activity. While such assays are very sensitive and specific for endo-cellulase (exo-acting cellulase enzymes produce little or no change in viscosity), they are limited by the fact that it is hard to define activity in conventional enzyme units (micromoles of substrate hydrolyzed or product produced per minute).
Cellooligosaccharide substrates
The lower DP cello-oligosaccharides (DP2-6) are sufficiently soluble in water to act as viable substrates for cellulase enzymes.[21] However, as these substrates are themselves 'reducing sugars', they are not suitable for use in traditional reducing sugar assays because they generate a high 'blank' value. However their cellulase mediated hydrolysis can be monitored by HPLC or IC methods to gain valuable information on the substrate requirements of a particular cellulase enzyme.
Reduced cellooligosaccharide substrates
Cello-oligosaccharides can be chemically reduced through the action of sodium borohydride to produce their corresponding sugar alcohols. These compounds do not react in reducing sugar assays but their hydrolysis products do. This makes borohydride reduced cello-oligosaccharides valuable substrates for the assay of cellulase using traditional reducing sugar assays such as the Nelson-Symogyi method.[22][23]
Dyed polysaccharide substrates[24]
These substrates can be subdivided into two classes-
- Insoluble chromogenic substrates: An insoluble cellulase substrate such as AZCL-HE-cellulose absorbs water to create gelatinous particles when placed in solution. This substrate is gradually depolymerised and solubilised by the action of cellulase. The reaction is terminated by adding an alkaline solution to stop enzyme activity and the reaction slurry is filtered or centrifuged. The colour in the filtrate or supernatant is measured and can be related to enzyme activity.
- Soluble chromogenic substrates: A cellulase sample is incubated with a water-soluble substrate such as azo-CM-cellulose, the reaction is terminated and high molecular weight, partially hydrolysed fragments are precipitated from solution with an organic solvent such as ethanol or methoxyethanol. The suspension is mixed thoroughly, centrifuged, and the colour in the supernatant solution (due to small, soluble, dyed fragments) is measured. With the aid of a standard curve, the enzyme activity can be determined.
Enzyme coupled reagents
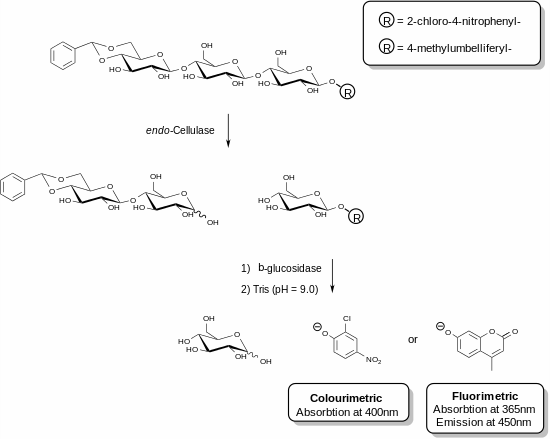
Recently, new reagents have been developed that allow for the specific measurement of endo-cellulase.[25][26] These methods involve the use of functionalised oligosaccharide substrates in the presence of an ancillary enzyme. In the example shown, a cellulase enzyme is able to recognise the trisaccharide fragment of cellulose and cleave this unit. The ancillary enzyme present in the reagent mixture (β-glucosidase) then acts to hydrolyse the fragment containing the chromophore or fluorophore. The assay is terminated by the addition of a basic solution that stops the enzymatic reaction and deprotonates the liberated phenolic compound to produce the phenolate species. The cellulase activity of a given sample is directly proportional to the quantity of phenolate liberated which can be measured using a spectrophotometer. The acetal functionalisation on the non-reducing end of the trisaccharide substrate prevents the action of the ancillary β-glucosidase on the parent substrate.
See also
- Cellulose 1,4-beta-cellobiosidase, an efficient cellulase
- Cellulase unit, a unit for quantifying cellulase activity
References
- PDB: 1NLR; Sulzenbacher G, Shareck F, Morosoli R, Dupont C, Davies GJ (December 1997). "The Streptomyces lividans family 12 endoglucanase: construction of the catalytic core, expression, and X-ray structure at 1.75 Å resolution". Biochemistry. 36 (51): 16032–9. doi:10.1021/bi972407v. PMID 9440876.; rendered with PyMOL
- Barkalow DG, Whistler RL. "Cellulose". AccessScience, McGraw-Hill.
- Bignell DE, Roisin Y, Lo N (2011). Biology of termites: a modern synthesis. Dordrecht: Springer. ISBN 978-9048139767.
- Watanabe H, Noda H, Tokuda G, Lo N (July 1998). "A cellulase gene of termite origin". Nature. 394 (6691): 330–1. Bibcode:1998Natur.394..330W. doi:10.1038/28527. PMID 9690469.
- Watanabe H, Tokuda G (August 2001). "Animal cellulases". Cellular and Molecular Life Sciences. 58 (9): 1167–78. doi:10.1007/PL00000931. PMID 11577976.
- Guerriero G, Sergeant K, Legay S. Hausman J-F, Cauchie H-M, Ahmad I, Siddiqui KS. 2018 Novel insights from comparative in silico analysis of green microalgae cellulases. Int. J. Mol. Sci. 19 (6), 1782.
- Zverlov VV, Schantz N, Schwarz WH (August 2005). "A major new component in the cellulosome of Clostridium thermocellum is a processive endo-beta-1,4-glucanase producing cellotetraose". FEMS Microbiology Letters. 249 (2): 353–8. doi:10.1016/j.femsle.2005.06.037. PMID 16006068.
- Payne CM, Bomble YJ, Taylor CB, McCabe C, Himmel ME, Crowley MF, Beckham GT (November 2011). "Multiple functions of aromatic-carbohydrate interactions in a processive cellulase examined with molecular simulation". The Journal of Biological Chemistry. 286 (47): 41028–35. doi:10.1074/jbc.M111.297713. PMC 3220501. PMID 21965672.
- Lee YJ, Kim BK, Lee BH, Jo KI, Lee NK, Chung CH, et al. (January 2008). "Purification and characterization of cellulase produced by Bacillus amyoliquefaciens DL-3 utilizing rice hull". Bioresource Technology. 99 (2): 378–86. doi:10.1016/j.biortech.2006.12.013. PMID 17320379.
- Cheng YS, Ko TP, Wu TH, Ma Y, Huang CH, Lai HL, et al. (April 2011). "Crystal structure and substrate-binding mode of cellulase 12A from Thermotoga maritima". Proteins. 79 (4): 1193–204. doi:10.1002/prot.22953. PMID 21268113.
- Liu Y, Yoshida M, Kurakata Y, Miyazaki T, Igarashi K, Samejima M, et al. (March 2010). "Crystal structure of a glycoside hydrolase family 6 enzyme, CcCel6C, a cellulase constitutively produced by Coprinopsis cinerea". The FEBS Journal. 277 (6): 1532–42. doi:10.1111/j.1742-4658.2010.07582.x. PMID 20148970.
- Tsai SL, DaSilva NA, Chen W (January 2013). "Functional display of complex cellulosomes on the yeast surface via adaptive assembly". ACS Synthetic Biology. 2 (1): 14–21. CiteSeerX 10.1.1.701.5515. doi:10.1021/sb300047u. PMID 23656322.
- Ravachol J, Borne R, Tardif C, de Philip P, Fierobe HP (March 2014). "Characterization of all family-9 glycoside hydrolases synthesized by the cellulosome-producing bacterium Clostridium cellulolyticum". The Journal of Biological Chemistry. 289 (11): 7335–48. doi:10.1074/jbc.M113.545046. PMC 3953250. PMID 24451379.
- Worthington Biochemical Corporation (2014), Cellulase. Accessed on 2014-07-03
- Bayer EA, Chanzy H, Lamed R, Shoham Y (October 1998). "Cellulose, cellulases and cellulosomes". Current Opinion in Structural Biology. 8 (5): 548–57. doi:10.1016/S0959-440X(98)80143-7. PMID 9818257.
- Bhaumik, Prasenjit; Dhepe, Paresh Laxmikant (2015-01-01). "Chapter 1. Conversion of Biomass into Sugars". Biomass Sugars for Non-Fuel Applications. Green Chemistry Series. Royal Society of Chemistry. pp. 1–53. doi:10.1039/9781782622079-00001. ISBN 978-1-78262-113-3.
- Fleming D, Rumbaugh KP (April 2017). "Approaches to Dispersing Medical Biofilms". Microorganisms. 5 (2): 15. doi:10.3390/microorganisms5020015. PMC 5488086. PMID 28368320.
- Fleming D, Chahin L, Rumbaugh K (February 2017). "Glycoside Hydrolases Degrade Polymicrobial Bacterial Biofilms in Wounds". Antimicrobial Agents and Chemotherapy. 61 (2): AAC.01998–16. doi:10.1128/AAC.01998-16. PMC 5278739. PMID 27872074.
- Jasani H, Umretiya N, Dharajiya D, Kapuria M, Shah S, Patel J (June 2016). "Isolation, optimization and production of cellulase by Aspergillus niger from agricultural waste". Journal of Pure and Applied Microbiology. 10 (2): 1159–66.
- Umezurike GM (January 1979). "The cellulolytic enzymes of Botryodiplodia theobromae Pat. Separation and characterization of cellulases and beta-glucosidases". The Biochemical Journal. 177 (1): 9–19. doi:10.1042/bj1770009. PMC 1186335. PMID 106849.
- Telke AA, Zhuang N, Ghatge SS, Lee SH, Ali Shah A, Khan H, et al. (2013). "Engineering of family-5 glycoside hydrolase (Cel5A) from an uncultured bacterium for efficient hydrolysis of cellulosic substrates". PLOS One. 8 (6): e65727. Bibcode:2013PLoSO...865727T. doi:10.1371/journal.pone.0065727. PMC 3681849. PMID 23785445.
- Nelson N (1944). "A photometric adaptation of the Somogyi method for the determination of glucose". J. Biol. Chem. 153: 375–80.
- Smogyi M (March 1952). "Notes on sugar determination". The Journal of Biological Chemistry. 195 (1): 19–23. PMID 14938350.
- McCleary BV (November 1980). "New chromogenic substrates for the assay of alpha-amylase and (1 leads to 4)-beta-D-glucanase". Carbohydrate Research. 86 (1): 97–104. doi:10.1016/s0008-6215(00)84584-x. PMID 6159974.
- McCleary BV, Mangan D, Daly R, Fort S, Ivory R, McCormack N (February 2014). "Novel substrates for the measurement of endo-1,4-β-glucanase (endo-cellulase)". Carbohydrate Research. 385: 9–17. doi:10.1016/j.carres.2013.12.001. PMID 24398300.
- Mangan D, McCleary BV, Liadova A, Ivory R, McCormack N (August 2014). "Quantitative fluorometric assay for the measurement of endo-1,4-β-glucanase". Carbohydrate Research. 395: 47–51. doi:10.1016/j.carres.2014.05.002. PMID 25038461.
Further reading
- Chapin FS, Matson PA, Mooney HA (2002). Principles of terrestrial ecosystem ecology (PDF). New York: Springer. ISBN 978-0-387-95439-4. Archived from the original (PDF) on 2016-03-05. Retrieved 2014-07-04.
- The Merck Manual of Diagnosis and Therapy, Chapter 24
- Deka D, Bhargavi P, Sharma A, Goyal D, Jawed M, Goyal A (2011). "Enhancement of Cellulase Activity from a New Strain of Bacillus subtilis by Medium Optimization and Analysis with Various Cellulosic Substrates". Enzyme Research. 2011: 151656. doi:10.4061/2011/151656. PMC 3102325. PMID 21637325.
- Zafar M, Ahmed S, Khan MI, Jamil A (May 2014). "Recombinant expression and characterization of a novel endoglucanase from Bacillus subtilis in Escherichia coli". Molecular Biology Reports. 41 (5): 3295–302. doi:10.1007/s11033-014-3192-8. PMID 24493451.