Personalized medicine
Personalized medicine, also referred to as precision medicine, is a medical model that separates people into different groups—with medical decisions, practices, interventions and/or products being tailored to the individual patient based on their predicted response or risk of disease.[1] The terms personalized medicine, precision medicine, stratified medicine and P4 medicine are used interchangeably to describe this concept[1][2] though some authors and organisations use these expressions separately to indicate particular nuances.[2]
| Part of a series on |
| Genetics |
|---|
 |
| Key components |
| History and topics |
| Research |
|
| Personalized medicine |
| Personalized medicine |
While the tailoring of treatment to patients dates back at least to the time of Hippocrates,[3] the term has risen in usage in recent years given the growth of new diagnostic and informatics approaches that provide understanding of the molecular basis of disease, particularly genomics. This provides a clear evidence base on which to stratify (group) related patients.[1][4][5]
Among 14 Grand Challenges for Engineering, initiative sponsored by National Academy of Engineering (NAE), personalized medicine has been identified as a key and prospective approach to “achieve optimal individual health decisions”, therefore overcoming the challenge of “Engineer better medicines”.[6][7]
Development of concept
In personalised medicine, diagnostic testing is often employed for selecting appropriate and optimal therapies based on the context of a patient's genetic content or other molecular or cellular analysis.[8] The use of genetic information has played a major role in certain aspects of personalized medicine (e.g. pharmacogenomics), and the term was first coined in the context of genetics, though it has since broadened to encompass all sorts of personalization measures,[8] including the use of proteomics,[9] imaging analysis, nanoparticle-based theranostics,[10] among others.
Background
Basics
Every person has a unique variation of the human genome.[11] Although most of the variation between individuals has no effect on health, an individual's health stems from genetic variation with behaviors and influences from the environment.[12][13]
Modern advances in personalized medicine rely on technology that confirms a patient's fundamental biology, DNA, RNA, or protein, which ultimately leads to confirming disease. For example, personalised techniques such as genome sequencing can reveal mutations in DNA that influence diseases ranging from cystic fibrosis to cancer. Another method, called RNA-seq, can show which RNA molecules are involved with specific diseases. Unlike DNA, levels of RNA can change in response to the environment. Therefore, sequencing RNA can provide a broader understanding of a person's state of health. Recent studies have linked genetic differences between individuals to RNA expression,[14] translation,[15] and protein levels.[16]
The concepts of personalised medicine can be applied to new and transformative approaches to health care. Personalised health care is based on the dynamics of systems biology and uses predictive tools to evaluate health risks and to design personalised health plans to help patients mitigate risks, prevent disease and to treat it with precision when it occurs. The concepts of personalised health care are receiving increasing acceptance with the Veterans Administration committing to personalised, proactive patient driven care for all veterans.[17] In some instances personalised health care can be tailored to the markup of the disease causing agent instead of the patient's genetic markup; examples are drug resistant bacteria or viruses.[18]
Method
In order for physicians to know if a mutation is connected to a certain disease, researchers often do a study called a “genome-wide association study” (GWAS). A GWAS study will look at one disease, and then sequence the genome of many patients with that particular disease to look for shared mutations in the genome. Mutations that are determined to be related to a disease by a GWAS study can then be used to diagnose that disease in future patients, by looking at their genome sequence to find that same mutation. The first GWAS, conducted in 2005, studied patients with age-related macular degeneration (ARMD).[19] It found two different mutations, each containing only a variation in only one nucleotide (called single nucleotide polymorphisms, or SNPs), which were associated with ARMD. GWAS studies like this have been very successful in identifying common genetic variations associated with diseases. As of early 2014, over 1,300 GWAS studies have been completed.[20]
Disease risk assessment
Multiple genes collectively influence the likelihood of developing many common and complex diseases.[12] Personalised medicine can also be used to predict a person's risk for a particular disease, based on one or even several genes. This approach uses the same sequencing technology to focus on the evaluation of disease risk, allowing the physician to initiate preventive treatment before the disease presents itself in their patient. For example, if it is found that a DNA mutation increases a person's risk of developing Type 2 Diabetes, this individual can begin lifestyle changes that will lessen their chances of developing Type 2 Diabetes later in life.
Applications
Advances in personalised medicine will create a more unified treatment approach specific to the individual and their genome. Personalised medicine may provide better diagnoses with earlier intervention, and more efficient drug development and more targeted therapies.[21]
Diagnosis and intervention
Having the ability to look at a patient on an individual basis will allow for a more accurate diagnosis and specific treatment plan. Genotyping is the process of obtaining an individual's DNA sequence by using biological assays.[22] By having a detailed account of an individual's DNA sequence, their genome can then be compared to a reference genome, like that of the Human Genome Project, to assess the existing genetic variations that can account for possible diseases. A number of private companies, such as 23andMe, Navigenics, and Illumina, have created Direct-to-Consumer genome sequencing accessible to the public.[11] Having this information from individuals can then be applied to effectively treat them. An individual's genetic make-up also plays a large role in how well they respond to a certain treatment, and therefore, knowing their genetic content can change the type of treatment they receive.
An aspect of this is pharmacogenomics, which uses an individual's genome to provide a more informed and tailored drug prescription.[23] Often, drugs are prescribed with the idea that it will work relatively the same for everyone, but in the application of drugs, there are a number of factors that must be considered. The detailed account of genetic information from the individual will help prevent adverse events, allow for appropriate dosages, and create maximum efficacy with drug prescriptions.[11] For instance, warfarin is the FDA approved oral anticoagulant commonly prescribed to patients with blood clots. Due to warfarin’s significant interindividual variability in pharmacokinetics and pharmacodynamics, its rate of adverse events is among the highest of all commonly prescribed drugs.[6] However, with the discovery of polymorphic variants in CYP2C9 and VKORC1 genotypes, two genes that encode the individual anticoagulant response,[24][25] physicians can use patients’ gene profile to prescribe optimum doses of warfarin to prevent side effects such as major bleeding and to allow sooner and better therapeutic efficacy.[6] The pharmacogenomic process for discovery of genetic variants that predict adverse events to a specific drug has been termed toxgnostics.[26]
An aspect of a theranostic platform applied to personalized medicine can be the use of diagnostic tests to guide therapy. The tests may involve medical imaging such as MRI contrast agents (T1 and T2 agents), fluorescent markers (organic dyes and inorganic quantum dots), and nuclear imaging agents (PET radiotracers or SPECT agents).[27][28] or in vitro lab test[29] including DNA sequencing[30] and often involve deep learning algorithms that weigh the result of testing for several biomarkers.[31]
In addition to specific treatment, personalised medicine can greatly aid the advancements of preventive care. For instance, many women are already being genotyped for certain mutations in the BRCA1 and BRCA2 gene if they are predisposed because of a family history of breast cancer or ovarian cancer.[32] As more causes of diseases are mapped out according to mutations that exist within a genome, the easier they can be identified in an individual. Measures can then be taken to prevent a disease from developing. Even if mutations were found within a genome, having the details of their DNA can reduce the impact or delay the onset of certain diseases.[21] Having the genetic content of an individual will allow better guided decisions in determining the source of the disease and thus treating it or preventing its progression. This will be extremely useful for diseases like Alzheimer’s or cancers that are thought to be linked to certain mutations in our DNA.[21]
A tool that is being used now to test efficacy and safety of a drug specific to a targeted patient group/sub-group is companion diagnostics. This technology is an assay that is developed during or after a drug is made available on the market and is helpful in enhancing the therapeutic treatment available based on the individual.[33] These companion diagnostics have incorporated the pharmacogenomic information related to the drug into their prescription label in an effort to assist in making the most optimal treatment decision possible for the patient.[33]
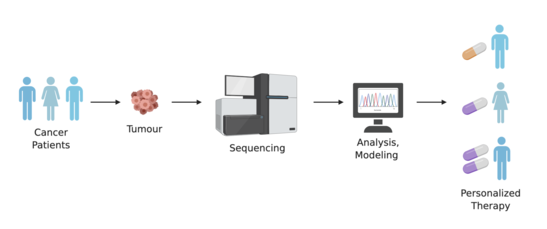
Drug development and usage
Having an individual's genomic information can be significant in the process of developing drugs as they await approval from the FDA for public use. Having a detailed account of an individual's genetic make-up can be a major asset in deciding if a patient can be chosen for inclusion or exclusion in the final stages of a clinical trial.[21] Being able to identify patients who will benefit most from a clinical trial will increase the safety of patients from adverse outcomes caused by the product in testing, and will allow smaller and faster trials that lead to lower overall costs.[34] In addition, drugs that are deemed ineffective for the larger population can gain approval by the FDA by using personal genomes to qualify the effectiveness and need for that specific drug or therapy even though it may only be needed by a small percentage of the population.,[21][35]
Today in medicine, it is common that physicians often use a trial and error strategy until they find the treatment therapy that is most effective for their patient.[21] With personalised medicine, these treatments can be more specifically tailored to an individual and give insight into how their body will respond to the drug and if that drug will work based on their genome.[11] The personal genotype can allow physicians to have more detailed information that will guide them in their decision in treatment prescriptions, which will be more cost-effective and accurate.[21] As quoted from the article Pharmacogenomics: The Promise of Personalised Medicine, “therapy with the right drug at the right dose in the right patient” is a description of how personalized medicine will affect the future of treatment.[36] For instance, Tamoxifen used to be a drug commonly prescribed to women with ER+ breast cancer, but 65% of women initially taking it developed resistance. After some research by people such as David Flockhart, it was discovered that women with certain mutation in their CYP2D6 gene, a gene that encodes the metabolizing enzyme, were not able to efficiently break down Tamoxifen, making it an ineffective treatment for their cancer.[37] Since then, women are now genotyped for those specific mutations, so that immediately these women can have the most effective treatment therapy.
Screening for these mutations is carried out via high-throughput screening or phenotypic screening. Several drug discovery and pharmaceutical companies are currently utilizing these technologies to not only advance the study of personalised medicine, but also to amplify genetic research; these companies include Alacris Theranostics, Persomics, Flatiron Health, Novartis, OncoDNA and Foundation Medicine, among others. Alternative multi-target approaches to the traditional approach of "forward" transfection library screening can entail reverse transfection or chemogenomics.
Pharmacy compounding is yet another application of personalised medicine. Though not necessarily utilizing genetic information, the customized production of a drug whose various properties (e.g. dose level, ingredient selection, route of administration, etc.) are selected and crafted for an individual patient is accepted as an area of personalised medicine (in contrast to mass-produced unit doses or fixed-dose combinations).
New methods are needed for delivering personalized drugs generated from pharmacy compounding efficiently to the disease sites of the body.[7] For instance, researchers are now trying to engineer nanocarriers that can precisely target the specific site by using real-time imaging and analyzing the pharmacodynamics of the drug delivery.[38] Currently, several candidate nanocarriers are being investigated, which are Iron oxide nanoparticles, quantum dots, carbon nanotubes, gold nanoparticles, and silica nanoparticles.[10] Alteration of surface chemistry allows these nanoparticles to be loaded with drugs, as well as to avoid the body’s immune response, making nanoparticle-based theranostics possible.[7][10] Nanocarriers’ targeting strategies are varied according to the disease. For example, if the disease is cancer, a common approach is to identify the biomarker expressed on the surface of cancer cells and to load its associated targeting vector onto nanocarrier to achieve recognition and binding; the size scale of the nanocarriers will also be engineered to reach the enhanced permeability and retention effect (EPR) in tumor targeting.[10] If the disease is localized in the specific organ, such as the kidney, the surface of the nanocarriers can be coated with a certain ligand that binds to the receptors inside that organ to achieve organ-targeting drug delivery and avoid non-specific uptake.[39] Despite the great potential of this nanoparticle-based drug delivery system, the significant progress in the field is yet to be made, and the nanocarriers are still being investigated and modified to meet clinical standards.[10][38]
Respiratory proteomics
.jpg)
Respiratory diseases affect humanity globally, with chronic lung diseases (e.g., asthma, chronic obstructive pulmonary disease, idiopathic pulmonary fibrosis, among others) and lung cancer causing extensive morbidity and mortality. These conditions are highly heterogeneous and require an early diagnosis. However, initial symptoms are nonspecific, and the clinical diagnosis is made late frequently. Over the last few years, personalized medicine has emerged as a medical care approach that uses novel technology [40] aiming to personalize treatments according to the particular patient's medical needs. In specific, proteomics is used to analyze a series of protein expressions, instead of a single biomarker.[41] Proteins control the body’s biological activities including health and disease, so proteomics is helpful in early diagnosis. In the case of respiratory disease, proteomics analyzes several biological samples including serum, blood cells, bronchoalveolar lavage fluids (BAL), nasal lavage fluids (NLF), sputum, among others.[41] The identification and quantification of complete protein expression from these biological samples are conducted by mass spectrometry and advanced analytical techniques.[42] Respiratory proteomics has made significant progress in the development of personalized medicine for supporting health care in recent years. For example, in a study conducted by Lazzari et al. in 2012, the proteomics-based approach has made substantial improvement in identifying multiple biomarkers of lung cancer that can be used in tailoring personalized treatments for individual patients.[43] More and more studies have demonstrated the usefulness of proteomics to provide targeted therapies for respiratory disease.[41]
Cancer genomics
Over recent decades cancer research has discovered a great deal about the genetic variety of types of cancer that appear the same in traditional pathology. There has also been increasing awareness of tumour heterogeneity, or genetic diversity within a single tumour. Among other prospects, these discoveries raise the possibility of finding that drugs that have not given good results applied to a general population of cases may yet be successful for a proportion of cases with particular genetic profiles.
"Personalized Onco-genomics" is the application of personalized medicine to Cancer Genomics, or “oncogenomics”. High-throughput sequencing methods are used to characterize genes associated with cancer to better understand disease pathology and improve drug development. Oncogenomics is one of the most promising branches of genomics, particularly because of its implications in drug therapy. Examples of this include:
- Trastuzumab (trade names Herclon, Herceptin) is a monoclonal antibody drug that interferes with the HER2/neu receptor. Its main use is to treat certain breast cancers. This drug is only used if a patient's cancer is tested for over-expression of the HER2/neu receptor. Two tissue-typing tests are used to screen patients for possible benefit from Herceptin treatment. The tissue tests are immunohistochemistry(IHC) and Fluorescence In Situ Hybridization(FISH)[44] Only Her2+ patients will be treated with Herceptin therapy (trastuzumab)[45]
- Tyrosine kinase inhibitors such as imatinib (marketed as Gleevec) have been developed to treat chronic myeloid leukemia (CML), in which the BCR-ABL fusion gene (the product of a reciprocal translocation between chromosome 9 and chromosome 22) is present in >95% of cases and produces hyperactivated abl-driven protein signaling. These medications specifically inhibit the Ableson tyrosine kinase (ABL) protein and are thus a prime example of "rational drug design" based on knowledge of disease pathophysiology.[46]
- The FoundationOne CDx report produced by Foundation Medicine, which looks at genes in individual patients' tumor biopsies and recommends specific drugs
- High mutation burden is indicative of response to immunotherapy, and also specific patterns of mutations have been associated with previous exposure to cytotoxic cancer drugs.[47]
Population screening
A lot of molecular-scale information about patients can be easily obtained through the use of genomics (microarray), proteomics (tissue array), and imaging (fMRI, micro-CT) technologies. These so-called molecular biomarkers such as genetic mutations have proven to be very powerful in disease prognosis, such as cancer prognosis.[48][49][50] The main three areas of cancer prediction fall under cancer recurrence, cancer susceptibility and cancer survivability.[51] When the molecular scale information is combined with macro-scale clinical data such as patients’ tumor type and other risk factors, the prognosis is significantly improved.[51] Consequently, given the use of molecular biomarkers, especially genomics, cancer prognosis or prediction has become very effective, especially when screening a large population.[52] Essentially, population genomics screening can be used to identify people at risk for disease, which can assist in preventative efforts. Notable examples include:[52]
- the MyCode® Community Health Initiative by Geisinger Health System, in which over 227,000 patients consented to genomic sequencing and follow-up counseling
- the Estonian Genome Project, which sequenced 52,000 Estonians
Challenges
As personalised medicine is practiced more widely, a number of challenges arise. The current approaches to intellectual property rights, reimbursement policies, patient privacy, data biases and confidentiality as well as regulatory oversight will have to be redefined and restructured to accommodate the changes personalised medicine will bring to healthcare.[53] Furthermore, the analysis of acquired diagnostic data is a recent challenge of personalized medicine and its implementation.[54] For example, genetic data obtained from next-generation sequencing requires computer-intensive data processing prior to its analysis.[55] In the future, adequate tools will be required to accelerate the adoption of personalised medicine to further fields of medicine, which requires the interdisciplinary cooperation of experts from specific fields of research, such as medicine, clinical oncology, biology, and artificial intelligence.
Regulatory oversight
The FDA has already started to take initiatives to integrate personalised medicine into their regulatory policies. An FDA report in October 2013 entitled, “Paving the Way for Personalized Medicine: FDA’s role in a New Era of Medical Product Development,” in which they outlined steps they would have to take to integrate genetic and biomarker information for clinical use and drug development.[56] They determined that they would have to develop specific regulatory science standards, research methods, reference material and other tools in order to incorporate personalised medicine into their current regulatory practices. For example, they are working on a “genomic reference library” for regulatory agencies to compare and test the validity of different sequencing platforms in an effort to uphold reliability.[56] A major challenge for those regulating personalized medicine is a way to demonstrate its effectiveness relative to the current standard of care. The new technology must be assessed for both clinical and cost effectiveness, and as it stands, regulatory agencies have no standardized method.[57]
Intellectual property rights
As with any innovation in medicine, investment and interest in personalised medicine is influenced by intellectual property rights.[53] There has been a lot of controversy regarding patent protection for diagnostic tools, genes, and biomarkers.[58] In June 2013, the U.S Supreme Court ruled that natural occurring genes cannot be patented, while “synthetic DNA” that is edited or artificially- created can still be patented. The Patent Office is currently reviewing a number of issues related to patent laws for personalised medicine, such as whether “confirmatory” secondary genetic tests post initial diagnosis, can have full immunity from patent laws. Those who oppose patents argue that patents on DNA sequences are an impediment to ongoing research while proponents point to research exemption and stress that patents are necessary to entice and protect the financial investments required for commercial research and the development and advancement of services offered.[58]
Reimbursement policies
Reimbursement policies will have to be redefined to fit the changes that personalised medicine will bring to the healthcare system. Some of the factors that should be considered are the level of efficacy of various genetic tests in the general population, cost-effectiveness relative to benefits, how to deal with payment systems for extremely rare conditions, and how to redefine the insurance concept of “shared risk” to incorporate the effect of the newer concept of “individual risk factors".[53] The study, Barriers to the Use of Personalized Medicine in Breast Cancer, took two different diagnostic tests which are BRACAnalysis and Oncotype DX. These tests have over ten-day turnaround times which results in the tests failing and delays in treatments. Patients are not being reimbursed for these delays which results in tests not being ordered. Ultimately, this leads to patients having to pay out-of-pocket for treatments because insurance companies do not want to accept the risks involved.[59]
Patient privacy and confidentiality
Perhaps the most critical issue with the commercialization of personalised medicine is the protection of patients. One of the largest issues is the fear and potential consequences for patients who are predisposed after genetic testing or found to be non-responsive towards certain treatments. This includes the psychological effects on patients due to genetic testing results. The right of family members who do not directly consent is another issue, considering that genetic predispositions and risks are inheritable. The implications for certain ethnic groups and presence of a common allele would also have to be considered.[53]
Moreover, we could refer to the privacy issue at all layers of personalized medicine from discovery to treatment. One of the leading issues is the consent of the patients to have their information used in genetic testing algorithms primarily AI algorithms. The consent of the institution who is providing the data to be used is of prominent concern as well.[60] In 2008, the Genetic Information Nondiscrimination Act (GINA) was passed in an effort to minimize the fear of patients participating in genetic research by ensuring that their genetic information will not be misused by employers or insurers.[53] On February 19, 2015 FDA issued a press release titled: "FDA permits marketing of first direct-to-consumer genetic carrier test for Bloom syndrome.[8]
Data Biases
Data biases also play an integral role in personalized medicine. It is important to ensure that the sample of genes being tested come from different populations. This is to ensure that the samples do not exhibit the same human biases we use in decision making.[61]
Consequently, if the designed algorithms for personalized medicine are biased, then the outcome of the algorithm will also be biased because of the lack of genetic testing in certain populations.[62] For instance, the results from the Framingham Heart Study have led to biased outcomes of predicting the risk of cardiovascular disease. This is because the sample was tested only on white people and when applied to the non-white population, the results were biased with overestimation and underestimation risks of cardiovascular disease.[63]
Implementation
Aside from issues related to the healthcare system, there are still several issues that must be addressed before personalized medicine can be implemented. Currently, very little of the human genome has been analyzed, and even if healthcare providers had access to a patient’s full genetic information, very little of it could be effectively leveraged into treatment.[64] Challenges also arise when processing such large amounts of genetic data. Even with error rates as low as 1 per 100 kb, processing a human genome could have roughly 30,000 errors.[65] This many errors, especially when trying to identify specific markers, can make discoveries, as well as verifiability difficult. There are methods to overcome this, but as it stands, they are computationally taxing, as well as expensive. There are also issues from an effectiveness standpoint, as after the genome has been processed, function in the variations among genomes must be analyzed using GWASs. While the impact of the SNPs discovered in these kinds of studies can be predicted, more work must be done to control for the vast amounts of variation that can occur because of the size of the genome being studied.[65] In order to effectively move forward in this area, steps must be taken to ensure the data being analyzed is good, and a wider view must be taken in terms of analyzing multiple SNPs for a phenotype. The most pressing issue that the implementation of personalized medicine is to apply the results of genetic mapping to improve the healthcare system. This is not only due to the infrastructure and technology required for a centralized database of genome data, but also the physicians that would have access to these tools would likely be unable to fully take advantage of them.[65] In order to truly implement a personalized medicine healthcare system, there must be an end-to-end change.
See also
References
- Stratified, personalised or P4 medicine: a new direction for placing the patient at the centre of healthcare and health education (Technical report). Academy of Medical Sciences. May 2015. Archived from the original on 27 October 2016. Retrieved 6 January 2016.
- "Many names for one concept or many concepts in one name?". PHG Foundation. Retrieved 6 January 2015.
- Egnew TR (1 March 2009). "Suffering, meaning, and healing: challenges of contemporary medicine". Annals of Family Medicine. 7 (2): 170–5. doi:10.1370/afm.943. PMC 2653974. PMID 19273873.
- "The Case for Personalized Medicine" (PDF). Personalized Medicine Coalition. 2014. Retrieved 6 January 2016.
- Smith, Richard (15 October 2012). "Stratified, personalised, or precision medicine". British Medical Journal. Retrieved 6 January 2016.
- Lesko LJ (June 2007). "Personalized medicine: elusive dream or imminent reality?". Clinical Pharmacology and Therapeutics. 81 (6): 807–16. doi:10.1038/sj.clpt.6100204. PMID 17505496.
- "Grand Challenges - Engineer Better Medicines". www.engineeringchallenges.org. Retrieved 2020-08-03.
- "Personalized Medicine 101". Personalized Medicine Coalition. Retrieved 26 April 2014.
- Priyadharshini, V. S.; Teran, Luis M. (2016-01-01), Donev, Rossen (ed.), "Chapter Five - Personalized Medicine in Respiratory Disease: Role of Proteomics", Advances in Protein Chemistry and Structural Biology, Personalized Medicine, Academic Press, 102, pp. 115–146, retrieved 2020-08-03
- Xie J, Lee S, Chen X (August 2010). "Nanoparticle-based theranostic agents". Advanced Drug Delivery Reviews. 62 (11): 1064–79. doi:10.1016/j.addr.2010.07.009. PMC 2988080. PMID 20691229.
- Dudley, J; Karczewski, K. (2014). Exploring Personal Genomics. Oxford: Oxford University Press.
- "Personalized Medicine 101: The Science". Personalized Medicine Coalition. Retrieved 26 April 2014.
- Lu YF, Goldstein DB, Angrist M, Cavalleri G (July 2014). "Personalized medicine and human genetic diversity". Cold Spring Harbor Perspectives in Medicine. 4 (9): a008581. doi:10.1101/cshperspect.a008581. PMC 4143101. PMID 25059740.
- Battle A, Mostafavi S, Zhu X, Potash JB, Weissman MM, McCormick C, et al. (January 2014). "Characterizing the genetic basis of transcriptome diversity through RNA-sequencing of 922 individuals". Genome Research. 24 (1): 14–24. doi:10.1101/gr.155192.113. PMC 3875855. PMID 24092820.
- Cenik C, Cenik ES, Byeon GW, Grubert F, Candille SI, Spacek D, et al. (November 2015). "Integrative analysis of RNA, translation, and protein levels reveals distinct regulatory variation across humans". Genome Research. 25 (11): 1610–21. doi:10.1101/gr.193342.115. PMC 4617958. PMID 26297486.
- Wu L, Candille SI, Choi Y, Xie D, Jiang L, Li-Pook-Than J, et al. (July 2013). "Variation and genetic control of protein abundance in humans". Nature. 499 (7456): 79–82. Bibcode:2013Natur.499...79W. doi:10.1038/nature12223. PMC 3789121. PMID 23676674.
- Snyderman, R. Personalized Health Care from Theory to Practice, Biotechnology J. 2012, 7
- Altmann A, Beerenwinkel N, Sing T, Savenkov I, Doumer M, Kaiser R, et al. (2007). "Improved prediction of response to antiretroviral combination therapy using the genetic barrier to drug resistance". Antiviral Therapy. 12 (2): 169–78. PMID 17503659.
- Haines JL, Hauser MA, Schmidt S, Scott WK, Olson LM, Gallins P, et al. (April 2005). "Complement factor H variant increases the risk of age-related macular degeneration". Science. 308 (5720): 419–21. Bibcode:2005Sci...308..419H. doi:10.1126/science.1110359. PMID 15761120. S2CID 32716116.
- "A Catalog of Published Genome-Wide Association Studies". Retrieved 28 June 2015.
- "Personalized Medicine 101: The Promise". Personalized Medicine Coalition. Retrieved April 26, 2014.
- "Research Portfolio Online Reporting Tools: Human Genome Project". National Institutes of Health (NIH). Retrieved April 28, 2014.
- "Genetics Home Reference: What is pharmacogenomics?". National Institutes of Health (NIH). Retrieved April 28, 2014.
- Breckenridge A, Orme M, Wesseling H, Lewis RJ, Gibbons R (April 1974). "Pharmacokinetics and pharmacodynamics of the enantiomers of warfarin in man". Clinical Pharmacology and Therapeutics. 15 (4): 424–30. doi:10.1002/cpt1974154424. PMID 4821443.
- Rieder MJ, Reiner AP, Gage BF, Nickerson DA, Eby CS, McLeod HL, et al. (June 2005). "Effect of VKORC1 haplotypes on transcriptional regulation and warfarin dose". The New England Journal of Medicine. 352 (22): 2285–93. doi:10.1056/NEJMoa044503. PMID 15930419.
- Church D, Kerr R, Domingo E, Rosmarin D, Palles C, Maskell K, et al. (June 2014). "'Toxgnostics': an unmet need in cancer medicine". Nature Reviews. Cancer. 14 (6): 440–5. doi:10.1038/nrc3729. PMID 24827503. S2CID 21601547.
- Xie J, Lee S, Chen X (August 2010). "Nanoparticle-based theranostic agents". Advanced Drug Delivery Reviews. Development of Theranostic Agents that Co-Deliver Therapeutic and Imaging Agents. 62 (11): 1064–79. doi:10.1016/j.addr.2010.07.009. PMC 2988080. PMID 20691229.
- Kelkar SS, Reineke TM (October 2011). "Theranostics: combining imaging and therapy". Bioconjugate Chemistry. 22 (10): 1879–903. doi:10.1021/bc200151q. PMID 21830812.
- Perkovic MN, Erjavec GN, Strac DS, Uzun S, Kozumplik O, Pivac N (March 2017). "Theranostic Biomarkers for Schizophrenia". International Journal of Molecular Sciences. 18 (4): 733. doi:10.3390/ijms18040733. PMC 5412319. PMID 28358316.
- Kamps R, Brandão RD, Bosch BJ, Paulussen AD, Xanthoulea S, Blok MJ, Romano A (January 2017). "Next-Generation Sequencing in Oncology: Genetic Diagnosis, Risk Prediction and Cancer Classification". International Journal of Molecular Sciences. 18 (2): 308. doi:10.3390/ijms18020308. PMC 5343844. PMID 28146134.
- Yahata N, Kasai K, Kawato M (April 2017). "Computational neuroscience approach to biomarkers and treatments for mental disorders". Psychiatry and Clinical Neurosciences. 71 (4): 215–237. doi:10.1111/pcn.12502. PMID 28032396.
- "Fact Sheet: BRCA1 and BRCA2: Cancer and Genetic Testing". National Cancer Institute (NCI). Retrieved April 28, 2014.
- "BIOMARKER TOOLKIT: Companion Diagnostics" (PDF). Amgen. Archived from the original (PDF) on August 1, 2014. Retrieved May 2, 2014.
- "Paving the Way for Personalized Medicine: FDA's Role in a New Era of Medical Product Development" (PDF). Federal Drug Administration (FDA). Retrieved April 28, 2014.
- Hamburg MA, Collins FS (July 2010). "The path to personalized medicine". The New England Journal of Medicine. 363 (4): 301–4. doi:10.1056/nejmp1006304. PMID 20551152.
- Mancinelli L, Cronin M, Sadée W (2000). "Pharmacogenomics: the promise of personalized medicine". AAPS PharmSci. 2 (1): 29–41. doi:10.1208/ps020104. PMC 2750999. PMID 11741220.
- Ellsworth RE, Decewicz DJ, Shriver CD, Ellsworth DL (May 2010). "Breast cancer in the personal genomics era". Current Genomics. 11 (3): 146–61. doi:10.2174/138920210791110951. PMC 2878980. PMID 21037853.
- Soni A, Gowthamarajan K, Radhakrishnan A (March 2018). "Personalized Medicine and Customized Drug Delivery Systems: The New Trend of Drug Delivery and Disease Management". International Journal of Pharmaceutical Compounding. 22 (2): 108–121. PMID 29877858.
- Wang, Jonathan; Poon, Christopher; Chin, Deborah; Milkowski, Sarah; Lu, Vivian; Hallows, Kenneth R.; Chung, Eun Ji (2018-10-01). "Design and in vivo characterization of kidney-targeting multimodal micelles for renal drug delivery". Nano Research. 11 (10): 5584–5595. doi:10.1007/s12274-018-2100-2. ISSN 1998-0000. S2CID 104241223.
- Priyadharshini VS, Teran LM (2016). "Personalized Medicine in Respiratory Disease: Role of Proteomics". Advances in Protein Chemistry and Structural Biology. 102: 115–46. doi:10.1016/bs.apcsb.2015.11.008. PMID 26827604.
- Priyadharshini, V. S.; Teran, Luis M. (2020-01-01), Faintuch, Joel; Faintuch, Salomao (eds.), "Chapter 24 - Role of respiratory proteomics in precision medicine", Precision Medicine for Investigators, Practitioners and Providers, Academic Press, pp. 255–261, ISBN 978-0-12-819178-1, retrieved 2020-08-03
- Fujii K, Nakamura H, Nishimura T (April 2017). "Recent mass spectrometry-based proteomics for biomarker discovery in lung cancer, COPD, and asthma". Expert Review of Proteomics. 14 (4): 373–386. doi:10.1080/14789450.2017.1304215. PMID 28271730. S2CID 3640863.
- Lazzari C, Spreafico A, Bachi A, Roder H, Floriani I, Garavaglia D, et al. (January 2012). "Changes in plasma mass-spectral profile in course of treatment of non-small cell lung cancer patients with epidermal growth factor receptor tyrosine kinase inhibitors". Journal of Thoracic Oncology. 7 (1): 40–8. doi:10.1097/JTO.0b013e3182307f17. PMID 21964534. S2CID 205738875.
- Carney, Walter (2006). "HER2/neu Status is an Important Biomarker in Guiding Personalized HER2/neu Therapy" (PDF). Connection. 9: 25–27.
- Telli ML, Hunt SA, Carlson RW, Guardino AE (August 2007). "Trastuzumab-related cardiotoxicity: calling into question the concept of reversibility". Journal of Clinical Oncology. 25 (23): 3525–33. doi:10.1200/JCO.2007.11.0106. PMID 17687157.
- Saglio G, Morotti A, Mattioli G, Messa E, Giugliano E, Volpe G, et al. (December 2004). "Rational approaches to the design of therapeutics targeting molecular markers: the case of chronic myelogenous leukemia". Annals of the New York Academy of Sciences. 1028 (1): 423–31. Bibcode:2004NYASA1028..423S. doi:10.1196/annals.1322.050. PMID 15650267.
- Pleasance E, Titmuss E, Williamson L, et al. (2020). "Pan-cancer analysis of advanced patient tumours reveals interactions between therapy and genomic landscapes". Nature Cancer. 1 (4): 452–468. doi:10.1038/s43018-020-0050-6.
- Duffy MJ (July 2001). "Biochemical markers in breast cancer: which ones are clinically useful?". Clinical Biochemistry. 34 (5): 347–52. doi:10.1016/S0009-9120(00)00201-0. PMID 11522269.
- Piccart M, Lohrisch C, Di Leo A, Larsimont D (2001). "The predictive value of HER2 in breast cancer". Oncology. 61 Suppl 2 (Suppl. 2): 73–82. doi:10.1159/000055405. PMID 11694791. S2CID 23123767.
- Baldus SE, Engelmann K, Hanisch FG (2004-01-01). "MUC1 and the MUCs: a family of human mucins with impact in cancer biology". Critical Reviews in Clinical Laboratory Sciences. 41 (2): 189–231. doi:10.1080/10408360490452040. PMID 15270554. S2CID 46460945.
- Cruz JA, Wishart DS (February 2007). "Applications of machine learning in cancer prediction and prognosis". Cancer Informatics. 2: 59–77. doi:10.1177/117693510600200030. PMC 2675494. PMID 19458758.
- Williams MS (August 2019). "Early Lessons from the Implementation of Genomic Medicine Programs". Annual Review of Genomics and Human Genetics. 20 (1): 389–411. doi:10.1146/annurev-genom-083118-014924. PMID 30811224.
- "Personalized Medicine 101: The Challenges". Personalized Medicine Coalition. Retrieved April 26, 2014.
- Huser V, Sincan M, Cimino JJ (2014). "Developing genomic knowledge bases and databases to support clinical management: current perspectives". Pharmacogenomics and Personalized Medicine. 7: 275–83. doi:10.2147/PGPM.S49904. PMC 4175027. PMID 25276091.
- "Analyze Genomes: Motivation". Schapranow, Matthieu-P. Retrieved July 20, 2014.
- "Paving the Way for Personalized Medicine: FDA's Role in a New Era of Medical Product Development" (PDF). U.S Food and Drug Administration. Retrieved April 26, 2014.
- Frueh FW (2013-09-01). "Regulation, reimbursement, and the long road of implementation of personalized medicine--a perspective from the United States". Value in Health. Personalized Medicine and the Role of Health Economics and Outcomes Research: Applications, Emerging Trends, and Future Research. 16 (6 Suppl): S27-31. doi:10.1016/j.jval.2013.06.009. PMID 24034309.
- "Intellectual Property Issues Impacting the Future of Personalized Medicine". American Intellectual Property Law Association. Retrieved April 26, 2014.
- Weldon CB, Trosman JR, Gradishar WJ, Benson AB, Schink JC (July 2012). "Barriers to the use of personalized medicine in breast cancer". Journal of Oncology Practice. 8 (4): e24-31. doi:10.1200/jop.2011.000448. PMC 3396824. PMID 23180995.
- Vayena E, Blasimme A, Cohen IG (November 2018). "Machine learning in medicine: Addressing ethical challenges". PLOS Medicine. 15 (11): e1002689. doi:10.1371/journal.pmed.1002689. PMC 6219763. PMID 30399149.
- Char DS, Shah NH, Magnus D (March 2018). "Implementing Machine Learning in Health Care - Addressing Ethical Challenges". The New England Journal of Medicine. 378 (11): 981–983. doi:10.1056/NEJMp1714229. PMC 5962261. PMID 29539284.
- Chernew ME, Landrum MB (March 2018). "Targeted Supplemental Data Collection - Addressing the Quality-Measurement Conundrum". The New England Journal of Medicine. 378 (11): 979–981. doi:10.1056/NEJMp1713834. PMID 29539286.
- Gijsberts CM, Groenewegen KA, Hoefer IE, Eijkemans MJ, Asselbergs FW, Anderson TJ, et al. (2015-07-02). "Race/Ethnic Differences in the Associations of the Framingham Risk Factors with Carotid IMT and Cardiovascular Events". PLOS ONE. 10 (7): e0132321. Bibcode:2015PLoSO..1032321G. doi:10.1371/journal.pone.0132321. PMC 4489855. PMID 26134404.
- Yngvadottir B, Macarthur DG, Jin H, Tyler-Smith C (2009-09-02). "The promise and reality of personal genomics". Genome Biology. 10 (9): 237. doi:10.1186/gb-2009-10-9-237. PMC 2768970. PMID 19723346.
- Fernald GH, Capriotti E, Daneshjou R, Karczewski KJ, Altman RB (July 2011). "Bioinformatics challenges for personalized medicine". Bioinformatics. 27 (13): 1741–8. doi:10.1093/bioinformatics/btr295. PMC 3117361. PMID 21596790.