Chromosome 6
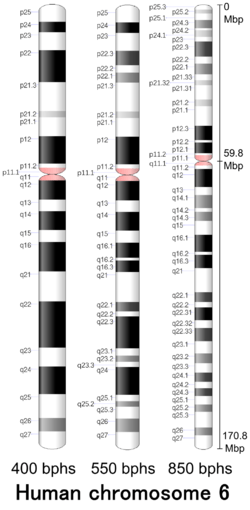
Chromosome 6 is one of the 23 pairs of chromosomes in humans. People normally have two copies of this chromosome. Chromosome 6 spans more than 170 million base pairs (the building material of DNA) and represents between 5.5 and 6% of the total DNA in cells. It contains the Major Histocompatibility Complex, which contains over 100 genes related to the immune response, and plays a vital role in organ transplantation.
| Chromosome 6 | |
|---|---|
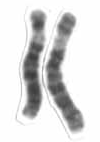 Human chromosome 6 pair after G-banding. One is from mother, one is from father. | |
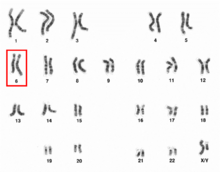 Chromosome 6 pair in human male karyogram. | |
| Features | |
| Length (bp) | 170,805,979 bp (GRCh38)[1] |
| No. of genes | 996 (CCDS)[2] |
| Type | Autosome |
| Centromere position | Submetacentric[3] (59.8 Mbp[4]) |
| Complete gene lists | |
| CCDS | Gene list |
| HGNC | Gene list |
| UniProt | Gene list |
| NCBI | Gene list |
| External map viewers | |
| Ensembl | Chromosome 6 |
| Entrez | Chromosome 6 |
| NCBI | Chromosome 6 |
| UCSC | Chromosome 6 |
| Full DNA sequences | |
| RefSeq | NC_000006 (FASTA) |
| GenBank | CM000668 (FASTA) |
Genes
The human leukocyte antigen lies on chromosome 6, with the exception of the gene for β2-microglobulin (which is located on chromosome 15), and encodes cell-surface antigen-presenting proteins among other functions.
Number of genes
In 2003, the entirety of chromosome 6 was manually annotated for proteins, resulting in the identification of 1,557 genes, and 633 pseudogenes.[5]
The following are some of the newer gene count estimates. Because researchers use different approaches to genome annotation their predictions of the number of genes on each chromosome varies (for technical details, see gene prediction). Among various projects, the collaborative consensus coding sequence project (CCDS) takes an extremely conservative strategy. So CCDS's gene number prediction represents a lower bound on the total number of human protein-coding genes.[6]
| Estimated by | Protein-coding genes | Non-coding RNA genes | Pseudogenes | Source | Release date |
|---|---|---|---|---|---|
| CCDS | 996 | — | — | [2] | 2016-09-08 |
| HGNC | 1,007 | 422 | 736 | [7] | 2017-05-12 |
| Ensembl | 1,038 | 985 | 800 | [8] | 2017-03-29 |
| UniProt | 1,111 | — | — | [9] | 2018-02-28 |
| NCBI | 1,053 | 1,188 | 911 | [10][11][12] | 2017-05-19 |
Gene list
The following is a partial list of genes on human chromosome 6. For complete list, see the link in the infobox on the right.
p-arm
The following are some of the genes located on p-arm (short arm) of human chromosome 6:
- ADTRP: encoding protein Androgen-dependent TFPI-regulating protein
- APOM: encoding protein Apolipoprotein M (6p21.33)
- TSBP1: encoding protein TSBP1 (6p21.32)
- C6orf62: chromosome 6 open reading frame 62 (6p22.3)
- C6orf89: chromosome 6 open reading frame 89 (6p21.2)
- CDKAL1: CDK5 regulatory subunit associated protein 1 like 1 (6p22.3)
- COL11A2: collagen, type XI, alpha 2(6p21.3)
- CYP21A2: cytochrome P450, family 21, subfamily A, polypeptide 2 (6p21.33)
- DHX16: DEAH-box helicase 16 (6p21.33)
- DOM3Z: Decapping exoribonuclease (6p21.33)
- DSP: Desmoplakin gene linked to cardiomyopathy (6p24.3)
- ELOVL5: ELOVL fatty acid elongase 5 (6p12.1)
- FBXO9: F-box protein 9 (6p12.1)
- G6B: Protein G6b (6p21.33)
- GCNT2: N-acetyllactosaminide beta-1,6-N-acetylglucosaminyl-transferase (6p24.3)
- GMDS: GDP-mannose 4,6-dehydratase (6p25.3)
- HCG4P11: HLA complex group 4 pseudogene 11
- HFE: hemochromatosis (6p22.2)
- HIST1H2AH: histone cluster 1 H2A family member h (6p22.1)
- HLA-A, HLA-B, HLA-C: major histocompatibility complex (MHC), class I, A, B, and C loci. (6p21.3)
- HLA-DQA1 and HLA-DQB1 form HLA-DQ heterodimer MHC class II, DQ: Celiac1, IDDM (6p21.3)
- HLA-DRA, HLA-DRB1, HLA-DRB3, HLA-DRB4, HLA-DRB5 forms HLA-DR, heterodimer MHC class II, DR (6p21.3)
- HLA-DPA1 and HLA-DPB1 forms HLA-DP, MHC class II, DP (6p21.3)
- HLA-Cw*06:02: gene variation related to psoriasis (6p21.3)
- LY6G6E encoding protein Lymphocyte antigen 6 complex, locus G6E (pseudogene) (6p21.33)
- LST1: leukocyte specific transcript 1 (6p21.33)
- MIR4640: microRNA 4640 (6p21.33)
- MLIP: muscular LMNA interaction protein (6p12.1)
- MRPS18B: mitochondrial ribosomal protein S18B (6p21.33)
- MUT: methylmalonyl Coenzyme A mutase (6p12.3)
- NHLRC1: NHL repeat containing E3 ubiquitin protein ligase 1 (6p22.3)
- NOL7: nucleolar protein 7 (6p23)
- NQO2: N-ribosyldihydronicotinamide:quinone reductase 2 (6p25.2)
- NRSN1: neurensin 1 (6p22.3)
- NUDT3: nudix hydrolase 3 (6p21.31)
- PFDN6: prefoldin subunit 6 (6p21.32)
- PHACTR1: phosphatase and actin regulator 1 (6p24.1)
- PKHD1: polycystic kidney and hepatic disease 1 (autosomal recessive) (6p21.2-p12)
- PRICKLE4: prickle planar cell polarity protein 4 (6p21.1)
- PRSS16: protease, serine 16 (6p22.1)
- PSMB8-AS1: PSMB8 antisense RNA 1 (head to head) (6p21.32)
- RIPOR2: RHO family interacting cell polarization regulator 2 (6p22.3)
- RPL10A: encoding protein 60S ribosomal protein L10a (6p21.31)
- SKIV2L: Ski2 like RNA helicase (6p21.33)
- SSR1: signal sequence receptor subunit 1 (6p24.3)
- TCF19: transcription factor 19 (6p21.33)
- TCP11: t-complex 11 (6p21.31)
- TJAP1: tight junction associated protein 1 (6p21.1)
- TP53COR1 encoding protein Tumor protein p53 pathway corepressor 1 (non-protein coding)
- TMEM151B: encoding protein Transmembrane protein 151B
- TNXB: tenascin XB (6p21.3)
- TRAM2: translocation associated membrane protein 2 (6p12.2)
- UBR2: ubiquitin protein ligase E3 component n-recognin 2 (6p21.2)
- UNC5CL: encoding protein Unc-5 homolog C (C. elegans)-like
- VEGF: vascular endothelial growth factor A (angiogenic growth factor) (6p21.1)
- VPS52: GARP complex subunit
- ZNF76: zinc finger protein 76 (6p21.31)
- ZNF193: zinc finger protein 193 (6p22.1)
- ZNRD1: zinc ribbon domain containing 1 (6p22.1)
q-arm
The following are some of the genes located on q-arm (long arm) of human chromosome 6:
- AIM1: encoding protein Absent in melanoma 1 protein (6q21)
- AIG1: encoding protein Androgen-induced protein 1 (6q24.2)
- AKIRIN2: akirin 2 (6q15)
- ARG1: arginase 1 (6q23.2)
- BCKDHB: branched-chain keto acid dehydrogenase E1, beta polypeptide (maple syrup urine disease) (6q14.1)
- BMIQ3: body mass index QTL 3
- TMEM242 encoding transmembrane protein TMEM242
- C6orf58: chromosome 6 open reading frame 58 (6q22.33)
- CFAP206: encoding protein Cilia And Flagella Associated Protein 206
- CMD1F: cardiomyopathy, dilated 1F
- CMD1K: cardiomyopathy, dilated 1K
- CNR1: cannabinoid 1 receptor (6q14-q15)[13]
- DFNB38: deafness, autosomal recessive 38
- DYX4: dyslexia susceptibility 4
- ECT2L: encoding protein Epithelial cell transforming sequence 2 oncogene-like
- ESR1: Estrogen receptor 1 (6q25)
- EYA4: eyes absent homolog 4 (Drosophila)(6q23.2)
- FBXL4: F-box and leucine rich repeat protein 4 (6q16.1-q16.2)
- FEB5: febrile convulsions 5
- HACE1: HECT domain and Ankyrin repeat containing, E3 ubiquitin protein ligase 1 (6q21)
- HEBP2: heme binding protein 2 (6q24.1)
- IDDM8: insulin dependent diabetes mellitus 8
- IDDM15: insulin dependent diabetes mellitus 15
- IFNGR: interferon-γ receptor gene (6q23-q24)
- IGF2R: insulin-like growth factor 2 receptor (6q25.3)
- IMPG1: interphotoreceptor matrix proteoglylcan 1 (6q14.1)
- LIN28B: lin-28 homolog B (6q16.3-q21)
- MAN1A1: mannosidase alpha class 1A member 1 (6q22.31)
- MB21D1: encoding protein Mab-21 domain containing 1
- MCDR1: macular dystrophy, retinal, 1
- MDN1: midasin AAA ATPase 1 (6q15)
- MOXD1: monooxygenase DBH like 1 (6q23.2)
- MTO1: mitochondrial tRNA translation optimization 1 (6q13)
- MRT18: mental retardation, non-syndromic, autosomal recessive
- MRT28: mental retardation, non-syndromic, autosomal recessive
- MTRF1L: mitochondrial translational release factor 1 like (6q25.2)
- MYO6: myosin VI (6q14.1)
- OA3: ocular albinism 3
- OPRM1: μ-opioid receptors (6q24-q25)
- OTSC7: otosclerosis 7
- PLG: plasminogen (6q26)
- PBCRA1
- PARK2: Parkinson disease (autosomal recessive, juvenile) 2, parkin (6q26)
- PCMT1: protein-L-isoaspartate (D-aspartate) O-methyltransferase (6q25.1)
- PERP: p53 apoptosis effector related to PMP-22 (6q23.3)
- PKIB: cAMP-dependent protein kinase inhibitor beta (6p22.31)
- PLAGL1: (6q24.2)
- RCD1: retinal cone dystrophy 1
- RP63: retinitis pigmentosa 63
- SASH1: SAM and SH3 domain containing 1 (6q24.3-q25.1)
- SCZD5: schizophrenia disorder 5
- SEN6: senescence (cellular)-related 6
- SENP6: SUMO1/sentrin specific peptidase 6 (6q14.1)
- SERAC1: serine active site containing 1 (6q25.3)
- SERINC1: serine incorporator 1 (6q22.31)
- SF3B5: splicing factor 3b subunit 5 (6q24.2)
- SMAP1: small ArfGAP 1 (6q13)
- SOBP: sine oculis binding protein homolog (6q21)
- SPG25: spastic paraplegia 25
- ST8:suppression of tumorigenicity 8
- SYNJ2: synaptojanin 2 (6q25.3)
- T: T brachyury transcription factor (more commonly known as the T gene) linked to Hepatocellular carcinoma and Chordoma (6q27)[14]
- TAAR1: trace amine associated receptor 1 (6q23.1)
- TAAR2: trace amine associated receptor 2 (6q24)
- TMEM200A: encoding protein Transmembrane protein 200A
- TSPYL1: TSPY like 1 (6q22.1)
- UNC93A: encoding protein Unc-93 homolog A (C. elegans)
- VNN1: vanin 1 (6q23.2)
- VNN2: vanin 2 (6q23.2)
- VTA1: Vesicle trafficking 1 (6q24.1-2)
- ZC2HC1: encoding protein Zinc finger C2HC-type containing 1B
- ZDHHC14: encoding protein Zinc finger, DHHC-type containing 14
Diseases and disorders
The following diseases are some of those related to genes on chromosome 6:
- ankylosing spondylitis, HLA-B
- collagenopathy, types II and XI
- Coeliac disease HLA-DQA1 & DQB1
- Ehlers-Danlos syndrome, classical, hypermobility, and Tenascin-X types
- Hashimoto's thyroiditis
- hemochromatosis
- Hemochromatosis type 1
- 21-hydroxylase deficiency
- maple syrup urine disease
- methylmalonic acidemia
- Autosomal nonsyndromic deafness
- otospondylomegaepiphyseal dysplasia
- Parkinson disease
- polycystic kidney disease
- porphyria
- porphyria cutanea tarda
- Rheumatoid arthritis, HLA-DR
- Stickler syndrome, COL11A2
- Systemic lupus erythematosus
- Diabetes mellitus type 1, HLA-DR, DQA1 & DQB1
- X-linked sideroblastic anemia
- Epilepsy
- Guillain Barre Syndrome
- Chordoma
- Hepatocellular carcinoma
- Schizophrenia
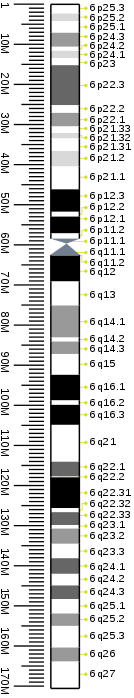
Cytogenetic band
| Chr. | Arm[20] | Band[21] | ISCN start[22] |
ISCN stop[22] |
Basepair start |
Basepair stop |
Stain[23] | Density |
|---|---|---|---|---|---|---|---|---|
| 6 | p | 25.3 | 0 | 118 | 1 | 2,300,000 | gneg | |
| 6 | p | 25.2 | 118 | 207 | 2,300,001 | 4,200,000 | gpos | 25 |
| 6 | p | 25.1 | 207 | 355 | 4,200,001 | 7,100,000 | gneg | |
| 6 | p | 24.3 | 355 | 548 | 7,100,001 | 10,600,000 | gpos | 50 |
| 6 | p | 24.2 | 548 | 592 | 10,600,001 | 11,600,000 | gneg | |
| 6 | p | 24.1 | 592 | 740 | 11,600,001 | 13,400,000 | gpos | 25 |
| 6 | p | 23 | 740 | 844 | 13,400,001 | 15,200,000 | gneg | |
| 6 | p | 22.3 | 844 | 1185 | 15,200,001 | 25,200,000 | gpos | 75 |
| 6 | p | 22.2 | 1185 | 1348 | 25,200,001 | 27,100,000 | gneg | |
| 6 | p | 22.1 | 1348 | 1585 | 27,100,001 | 30,500,000 | gpos | 50 |
| 6 | p | 21.33 | 1585 | 1718 | 30,500,001 | 32,100,000 | gneg | |
| 6 | p | 21.32 | 1718 | 1836 | 32,100,001 | 33,500,000 | gpos | 25 |
| 6 | p | 21.31 | 1836 | 2162 | 33,500,001 | 36,600,000 | gneg | |
| 6 | p | 21.2 | 2162 | 2310 | 36,600,001 | 40,500,000 | gpos | 25 |
| 6 | p | 21.1 | 2310 | 2755 | 40,500,001 | 46,200,000 | gneg | |
| 6 | p | 12.3 | 2755 | 3080 | 46,200,001 | 51,800,000 | gpos | 100 |
| 6 | p | 12.2 | 3080 | 3140 | 51,800,001 | 53,000,000 | gneg | |
| 6 | p | 12.1 | 3140 | 3377 | 5,300,0001 | 57,200,000 | gpos | 100 |
| 6 | p | 11.2 | 3377 | 3421 | 57,200,001 | 58,500,000 | gneg | |
| 6 | p | 11.1 | 3421 | 3554 | 58,500,001 | 59,800,000 | acen | |
| 6 | q | 11.1 | 3554 | 3658 | 59,800,001 | 62,600,000 | acen | |
| 6 | q | 11.2 | 3658 | 3732 | 62,600,001 | 62,700,000 | gneg | |
| 6 | q | 12 | 3732 | 4147 | 62,700,001 | 69,200,000 | gpos | 100 |
| 6 | q | 13 | 4147 | 4324 | 69,200,001 | 75,200,000 | gneg | |
| 6 | q | 14.1 | 4324 | 4621 | 75,200,001 | 83,200,000 | gpos | 50 |
| 6 | q | 14.2 | 4621 | 4709 | 83,200,001 | 84,200,000 | gneg | |
| 6 | q | 14.3 | 4709 | 4917 | 84,200,001 | 87,300,000 | gpos | 50 |
| 6 | q | 15 | 4917 | 5228 | 87,300,001 | 92,500,000 | gneg | |
| 6 | q | 16.1 | 5228 | 5613 | 92,500,001 | 98,900,000 | gpos | 100 |
| 6 | q | 16.2 | 5613 | 5687 | 98,900,001 | 100,000,000 | gneg | |
| 6 | q | 16.3 | 5687 | 5983 | 10,000,0001 | 105,000,000 | gpos | 100 |
| 6 | q | 21 | 5983 | 6531 | 10,500,0001 | 114,200,000 | gneg | |
| 6 | q | 22.1 | 6531 | 6753 | 114,200,001 | 117,900,000 | gpos | 75 |
| 6 | q | 22.2 | 6753 | 6872 | 117,900,001 | 118,100,000 | gneg | |
| 6 | q | 22.31 | 6872 | 7168 | 118,100,001 | 125,800,000 | gpos | 100 |
| 6 | q | 22.32 | 7168 | 7345 | 125,800,001 | 126,800,000 | gneg | |
| 6 | q | 22.33 | 7345 | 7642 | 126,800,001 | 130,000,000 | gpos | 75 |
| 6 | q | 23.1 | 7642 | 7923 | 13,000,0001 | 130,900,000 | gneg | |
| 6 | q | 23.2 | 7923 | 8145 | 130,900,001 | 134,700,000 | gpos | 50 |
| 6 | q | 23.3 | 8145 | 8352 | 134,700,001 | 138,300,000 | gneg | |
| 6 | q | 24.1 | 8352 | 8560 | 138,300,001 | 142,200,000 | gpos | 75 |
| 6 | q | 24.2 | 8560 | 8708 | 142,200,001 | 145,100,000 | gneg | |
| 6 | q | 24.3 | 8708 | 8886 | 145,100,001 | 148,500,000 | gpos | 75 |
| 6 | q | 25.1 | 8886 | 9078 | 148,500,001 | 152,100,000 | gneg | |
| 6 | q | 25.2 | 9078 | 9241 | 152,100,001 | 155,200,000 | gpos | 50 |
| 6 | q | 25.3 | 9241 | 9596 | 155,200,001 | 160,600,000 | gneg | |
| 6 | q | 26 | 9596 | 9774 | 160,600,001 | 164,100,000 | gpos | 50 |
| 6 | q | 27 | 9774 | 10100 | 164,100,001 | 170,805,979 | gneg |
References
- "Human Genome Assembly GRCh38 - Genome Reference Consortium". National Center for Biotechnology Information. 2013-12-24. Retrieved 2017-03-04.
- "Search results - 6[CHR] AND "Homo sapiens"[Organism] AND ("has ccds"[Properties] AND alive[prop]) - Gene". NCBI. CCDS Release 20 for Homo sapiens. 2016-09-08. Retrieved 2017-05-28.
- Tom Strachan; Andrew Read (2 April 2010). Human Molecular Genetics. Garland Science. p. 45. ISBN 978-1-136-84407-2.
- Genome Decoration Page, NCBI. Ideogram data for Homo sapience (850 bphs, Assembly GRCh38.p3). Last update 2014-06-03. Retrieved 2017-04-26.
- Mungall AJ, Palmer SA, Sims SK, Edwards CA, Ashurst JL, Wilming L, et al. (October 2003). "The DNA sequence and analysis of human chromosome 6". Nature. 425 (6960): 805–11. Bibcode:2003Natur.425..805M. doi:10.1038/nature02055. PMID 14574404.
- Pertea M, Salzberg SL (2010). "Between a chicken and a grape: estimating the number of human genes". Genome Biol. 11 (5): 206. doi:10.1186/gb-2010-11-5-206. PMC 2898077. PMID 20441615.
- "Statistics & Downloads for chromosome 6". HUGO Gene Nomenclature Committee. 2017-05-12. Retrieved 2017-05-19.
- "Chromosome 6: Chromosome summary - Homo sapiens". Ensembl Release 88. 2017-03-29. Retrieved 2017-05-19.
- "Human chromosome 6: entries, gene names and cross-references to MIM". UniProt. 2018-02-28. Retrieved 2018-03-16.
- "Search results - 6[CHR] AND "Homo sapiens"[Organism] AND ("genetype protein coding"[Properties] AND alive[prop]) - Gene". NCBI. 2017-05-19. Retrieved 2017-05-20.
- "Search results - 6[CHR] AND "Homo sapiens"[Organism] AND ( ("genetype miscrna"[Properties] OR "genetype ncrna"[Properties] OR "genetype rrna"[Properties] OR "genetype trna"[Properties] OR "genetype scrna"[Properties] OR "genetype snrna"[Properties] OR "genetype snorna"[Properties]) NOT "genetype protein coding"[Properties] AND alive[prop]) - Gene". NCBI. 2017-05-19. Retrieved 2017-05-20.
- "Search results - 6[CHR] AND "Homo sapiens"[Organism] AND ("genetype pseudo"[Properties] AND alive[prop]) - Gene". NCBI. 2017-05-19. Retrieved 2017-05-20.
- Matsuda LA, Lolait SJ, Brownstein MJ, Young AC, Bonner TI (August 1990). "Structure of a cannabinoid receptor and functional expression of the cloned cDNA". Nature. 346 (6284): 561–4. Bibcode:1990Natur.346..561M. doi:10.1038/346561a0. PMID 2165569.
- "T brachyury transcription factor". T - T brachyury transcription factor - Genetics Home Reference.
- Genome Decoration Page, NCBI. Ideogram data for Homo sapience (400 bphs, Assembly GRCh38.p3). Last update 2014-03-04. Retrieved 2017-04-26.
- Genome Decoration Page, NCBI. Ideogram data for Homo sapience (550 bphs, Assembly GRCh38.p3). Last update 2015-08-11. Retrieved 2017-04-26.
- International Standing Committee on Human Cytogenetic Nomenclature (2013). ISCN 2013: An International System for Human Cytogenetic Nomenclature (2013). Karger Medical and Scientific Publishers. ISBN 978-3-318-02253-7.
- Sethakulvichai, W.; Manitpornsut, S.; Wiboonrat, M.; Lilakiatsakun, W.; Assawamakin, A.; Tongsima, S. (2012). Estimation of band level resolutions of human chromosome images. In Computer Science and Software Engineering (JCSSE), 2012 International Joint Conference on. pp. 276–282. doi:10.1109/JCSSE.2012.6261965. ISBN 978-1-4673-1921-8.
- Genome Decoration Page, NCBI. Ideogram data for Homo sapience (850 bphs, Assembly GRCh38.p3). Last update 2014-06-03. Retrieved 2017-04-26.
- "p": Short arm; "q": Long arm.
- For cytogenetic banding nomenclature, see article locus.
- These values (ISCN start/stop) are based on the length of bands/ideograms from the ISCN book, An International System for Human Cytogenetic Nomenclature (2013). Arbitrary unit.
- gpos: Region which is positively stained by G banding, generally AT-rich and gene poor; gneg: Region which is negatively stained by G banding, generally CG-rich and gene rich; acen Centromere. var: Variable region; stalk: Stalk.
- Notes
- Some text in this article was taken from http://ghr.nlm.nih.gov/chromosome=6 (public domain)
Further reading
- Gilbert F (2002). "Chromosome 6". Genet Test. 6 (4): 341–58. doi:10.1089/10906570260471912. PMID 12537662.
External links
| Wikimedia Commons has media related to Human chromosome 6. |
- National Institutes of Health. "Chromosome 6". Genetics Home Reference. Retrieved 2017-05-06.
- "Chromosome 6". Human Genome Project Information Archive 1990–2003. Retrieved 2017-05-06.
- "Chromosome 6 Research Project". Parent-driven research for genotype-phenotype studies on chromosome 6 disorders. Retrieved 2017-06-17