3C-like protease
The 3C-like protease (3CLpro), formally known as C30 Endopeptidase, is the main protease found in coronaviruses. It cleaves the coronavirus polyprotein at eleven conserved sites. It is a cysteine protease and a member of the PA clan of proteases. It has a cysteine-histidine catalytic dyad at its active site and cleaves a Gln–(Ser/Ala/Gly) peptide bond.
| SARS coronavirus main proteinase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
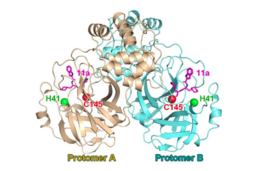 SARS-CoV-2 main proteinase dimer with the catalytic dyad (H41; C145) in complex with protease inhibitor chemical 11a | |||||||||
| Identifiers | |||||||||
| EC number | 3.4.22.69 | ||||||||
| Databases | |||||||||
| IntEnz | IntEnz view | ||||||||
| BRENDA | BRENDA entry | ||||||||
| ExPASy | NiceZyme view | ||||||||
| KEGG | KEGG entry | ||||||||
| MetaCyc | metabolic pathway | ||||||||
| PRIAM | profile | ||||||||
| PDB structures | RCSB PDB PDBe PDBsum | ||||||||
| |||||||||
| Peptidase C30, Coronavirus endopeptidase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Identifiers | |||||||||
| Symbol | Peptidase_C30 | ||||||||
| Pfam | PF05409 | ||||||||
| InterPro | IPR008740 | ||||||||
| PROSITE | PS51442 | ||||||||
| MEROPS | C30 | ||||||||
| SCOPe | d1q2wb1 / SUPFAM | ||||||||
| |||||||||
The Enzyme Commission refers to this family as SARS coronavirus main proteinase (Mpro; EC 3.4.22.69). The 3CL protease corresponds to coronavirus nonstructural protein 5 (nsp5). The "3C" in the common name refers to the 3C protease (3Cpro) which is a homologous protease found in picornaviruses.
Function
The 3C-like protease is able to catalytically cleave a peptide bond between a glutamine at position P1 and a small amino acid (serine, alanine, or glycine) at position P1'. The SARS coronavirus 3CLpro can for instance self-cleave the following peptides:[1][2][3]
TSAVLQ-SGFRK-NH2 and SGVTFQ-GKFKK are the two peptides corresponding to the two self-cleavage sites of the SARS 3C-like proteinase
The protease is important in the processing of the coronavirus replicase polyprotein (P0C6U8). It is the main protease in coronaviruses and corresponds to nonstructural protein 5 (nsp5).[4] It cleaves the coronavirus polyprotein at 11 conserved sites. The 3CL protease has a cysteine-histidine catalytic dyad at its active site.[2] The sulfur of the cysteine acts as a nucleophile and the imidazole ring of the histidine as a general base.[5]
Nomenclature
Alternative names provided by the EC include 3CLpro, 3C-like protease, coronavirus 3C-like protease, Mpro, SARS 3C-like protease, SARS coronavirus 3CL protease, SARS coronavirus main peptidase, SARS coronavirus main protease, SARS-CoV 3CLpro enzyme, SARS-CoV main protease, SARS-CoV Mpro and severe acute respiratory syndrome coronavirus main protease.
As a treatment target
The protease 3CLpro is a potential drug target for coronavirus infections due to its essential role in processing the polyproteins that are translated from the viral RNA. The X-ray structures of the unliganded SARS-CoV-2 protease 3CLpro and its complex with an α-ketoamide inhibitor provides a basis for design of α-ketoamide inhibitors[6] for a treatment of SARS-CoV-2 infection.[7][8][9][10][11] Potential protease inhibitors being developed against 3CLpro and homologous 3Cpro include CLpro-1, GC376, Rupintrivir, chemical 11a, and chemical 11b.[12][13][14][15]
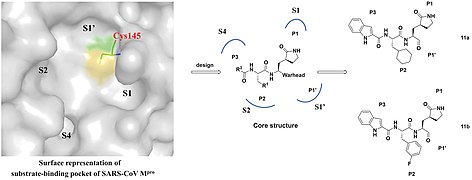 Design strategy of SARS-CoV-2 3CLpro inhibitors and the chemical structures of 11a and 11b.
Design strategy of SARS-CoV-2 3CLpro inhibitors and the chemical structures of 11a and 11b.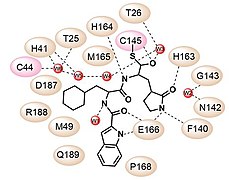 3CLpro inhibitor and chemical 11a
3CLpro inhibitor and chemical 11a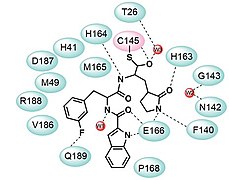 3CLpro inhibitor and chemical 11b
3CLpro inhibitor and chemical 11b
Other 3C(-like) proteases
3C-like proteases (3C(L)pro) are widely found in (+)ssRNA viruses. All of them are cysteine proteases with a chymotrypsin-like fold (PA clan), using a catalytic dyad or triad. They share some general similarities on substrate specificity and inhibitor effectiveness. They are divided into subfamilies by sequence similarity, corresponding to the family of viruses they are found in:[16]
- This entry is the coronavirus 3CLpro.
- Picornaviridae have a picronavirus 3Cpro (EC 3.4.22.28; InterPro: IPR000199; MEROPS C03). This is the earliest-studied family. Examples include the ones found in poliovirus and in rhinovirus (both are members of genus Enterovirus).
- Caliciviridae have a 3CLpro (InterPro: IPR001665; MEROPS C37). Examples include the one found in Norwalk virus.
Additional members are known from Potyviridae and non-Coronaviridae Nidovirales.[17]
References
- Goetz DH, Choe Y, Hansell E, Chen YT, McDowell M, Jonsson CB, Roush WR, McKerrow J, Craik CS (July 2007). "Substrate specificity profiling and identification of a new class of inhibitor for the major protease of the SARS coronavirus". Biochemistry. 46 (30): 8744–52. doi:10.1021/bi0621415. PMID 17605471.
- Fan K, Wei P, Feng Q, Chen S, Huang C, Ma L, Lai B, Pei J, Liu Y, Chen J, Lai L (January 2004). "Biosynthesis, purification, and substrate specificity of severe acute respiratory syndrome coronavirus 3C-like proteinase". The Journal of Biological Chemistry. 279 (3): 1637–42. doi:10.1074/jbc.m310875200. PMID 14561748.
- Akaji K, Konno H, Onozuka M, Makino A, Saito H, Nosaka K (November 2008). "Evaluation of peptide-aldehyde inhibitors using R188I mutant of SARS 3CL protease as a proteolysis-resistant mutant". Bioorganic & Medicinal Chemistry. 16 (21): 9400–8. doi:10.1016/j.bmc.2008.09.057. PMC 7126698. PMID 18845442.
- Fehr AR, Perlman S (2015). "Coronaviruses: an overview of their replication and pathogenesis". In Maier HJ, Bickerton E, Britton P (eds.). Coronaviruses. Methods in Molecular Biology. 1282. Springer. pp. 1–23. doi:10.1007/978-1-4939-2438-7_1. ISBN 978-1-4939-2438-7. PMC 4369385. PMID 25720466.
See section: Virion Structure.
- Ryu, Young Bae; Park, Su-Jin; Kim, Young Min; Lee, Ju-Yeon; Seo, Woo Duck; Chang, Jong Sun; Park, Ki Hun; Rho, Mun-Chual; Lee, Woo Song (2010-03-15). "SARS-CoV 3CLpro inhibitory effects of quinone-methide triterpenes from Tripterygium regelii". Bioorganic & Medicinal Chemistry Letters. 20 (6): 1873–1876. doi:10.1016/j.bmcl.2010.01.152. ISSN 0960-894X. PMC 7127101. PMID 20167482.
- Ocain TD, Rich DH (February 1992). "alpha-Keto amide inhibitors of aminopeptidases". Journal of Medicinal Chemistry. 35 (3): 451–6. doi:10.1021/jm00081a005. PMID 1738140.
- Anand K, Ziebuhr J, Wadhwani P, Mesters JR, Hilgenfeld R (June 2003). "Coronavirus main proteinase (3CLpro) structure: basis for design of anti-SARS drugs". Science. 300 (5626): 1763–7. Bibcode:2003Sci...300.1763A. doi:10.1126/science.1085658. PMID 12746549.
- Pacifico S, Ferretti V, Albanese V, Fantinati A, Gallerani E, Nicoli F, et al. (July 2019). "Synthesis and Biological Activity of Peptide α-Ketoamide Derivatives as Proteasome Inhibitors". ACS Medicinal Chemistry Letters. 10 (7): 1086–1092. doi:10.1021/acsmedchemlett.9b00233. PMC 6627721. PMID 31312413.
- Kusov Y, Tan J, Alvarez E, Enjuanes L, Hilgenfeld R (October 2015). "A G-quadruplex-binding macrodomain within the "SARS-unique domain" is essential for the activity of the SARS-coronavirus replication-transcription complex". Virology. 484: 313–22. doi:10.1016/j.virol.2015.06.016. PMC 4567502. PMID 26149721.
- Zhang L, Lin D, Kusov Y, Nian Y, Ma Q, Wang J, et al. (February 2020). "α-Ketoamides as Broad-Spectrum Inhibitors of Coronavirus and Enterovirus Replication: Structure-Based Design, Synthesis, and Activity Assessment". Journal of Medicinal Chemistry. 63 (9): 4562–4578. doi:10.1021/acs.jmedchem.9b01828. PMC 7098070. PMID 32045235.
- Zhang L, Lin D, Sun X, Curth U, Drosten C, Sauerhering L, et al. (March 2020). "Crystal structure of SARS-CoV-2 main protease provides a basis for design of improved α-ketoamide inhibitors". Science. 368 (6489): 409–412. Bibcode:2020Sci...368..409Z. doi:10.1126/science.abb3405. PMC 7164518. PMID 32198291.
- Morse JS, Lalonde T, Xu S, Liu WR (March 2020). "Learning from the Past: Possible Urgent Prevention and Treatment Options for Severe Acute Respiratory Infections Caused by 2019-nCoV". ChemBioChem. 21 (5): 730–738. doi:10.1002/cbic.202000047. PMC 7162020. PMID 32022370.
- Liu C, Zhou Q, Li Y, Garner LV, Watkins SP, Carter LJ, et al. (March 2020). "Research and Development on Therapeutic Agents and Vaccines for COVID-19 and Related Human Coronavirus Diseases". ACS Central Science. 6 (3): 315–331. doi:10.1021/acscentsci.0c00272. PMC 7094090. PMID 32226821.
- Ramajayam R, Tan KP, Liang PH (October 2011). "Recent development of 3C and 3CL protease inhibitors for anti-coronavirus and anti-picornavirus drug discovery". Biochemical Society Transactions. 39 (5): 1371–5. doi:10.1042/BST0391371. PMID 21936817.
- Dai, Wenhao; Zhang, Bing; Jiang, Xia-Ming; Su, Haixia; Li, Jian; Zhao, Yao; Xie, Xiong; Jin, Zhenming; Peng, Jingjing; Liu, Fengjiang; Li, Chunpu (2020-06-19). "Structure-based design of antiviral drug candidates targeting the SARS-CoV-2 main protease". Science. 368 (6497): 1331–1335. doi:10.1126/science.abb4489. ISSN 0036-8075. PMID 32321856.
- Kim Y, Lovell S, Tiew KC, Mandadapu SR, Alliston KR, Battaile KP, et al. (November 2012). "Broad-spectrum antivirals against 3C or 3C-like proteases of picornaviruses, noroviruses, and coronaviruses". Journal of Virology. 86 (21): 11754–62. doi:10.1128/JVI.01348-12. PMC 3486288. PMID 22915796.
- Ziebuhr J, Bayer S, Cowley JA, Gorbalenya AE (January 2003). "The 3C-like proteinase of an invertebrate nidovirus links coronavirus and potyvirus homologs". Journal of Virology. 77 (2): 1415–26. doi:10.1128/jvi.77.2.1415-1426.2003. PMC 140795. PMID 12502857.
Further reading
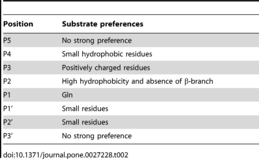
- Chuck CP, Chow HF, Wan DC, Wong KB (2 November 2011). "Profiling of substrate specificities of 3C-like proteases from group 1, 2a, 2b, and 3 coronaviruses". PLOS ONE. 6 (11): e27228. Bibcode:2011PLoSO...627228C. doi:10.1371/journal.pone.0027228. PMC 3206940. PMID 22073294.
External links
- SARS+coronavirus+main+proteinase at the US National Library of Medicine Medical Subject Headings (MeSH)
- Peptidase C30/C16 in coronavirus, InterPro: IPR013016. The MEROPS C16 one is the "papain-like" PL-PRO.