Chymotrypsin
Chymotrypsin (EC 3.4.21.1, chymotrypsins A and B, alpha-chymar ophth, avazyme, chymar, chymotest, enzeon, quimar, quimotrase, alpha-chymar, alpha-chymotrypsin A, alpha-chymotrypsin) is a digestive enzyme component of pancreatic juice acting in the duodenum, where it performs proteolysis, the breakdown of proteins and polypeptides.[2] Chymotrypsin preferentially cleaves peptide amide bonds where the side chain of the amino acid N-terminal to the scissile amide bond (the P1 position) is a large hydrophobic amino acid (tyrosine, tryptophan, and phenylalanine). These amino acids contain an aromatic ring in their side chain that fits into a hydrophobic pocket (the S1 position) of the enzyme. It is activated in the presence of trypsin. The hydrophobic and shape complementarity between the peptide substrate P1 side chain and the enzyme S1 binding cavity accounts for the substrate specificity of this enzyme.[3][4] Chymotrypsin also hydrolyzes other amide bonds in peptides at slower rates, particularly those containing leucine and methionine at the P1 position.
| Chymotrypsin | |||||||||
|---|---|---|---|---|---|---|---|---|---|
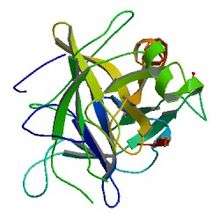 Crystallographic structure of Bos taurus chymotrypsinogen[1] | |||||||||
| Identifiers | |||||||||
| EC number | 3.4.21.1 | ||||||||
| CAS number | 9004-07-3 | ||||||||
| Databases | |||||||||
| IntEnz | IntEnz view | ||||||||
| BRENDA | BRENDA entry | ||||||||
| ExPASy | NiceZyme view | ||||||||
| KEGG | KEGG entry | ||||||||
| MetaCyc | metabolic pathway | ||||||||
| PRIAM | profile | ||||||||
| PDB structures | RCSB PDB PDBe PDBsum | ||||||||
| Gene Ontology | AmiGO / QuickGO | ||||||||
| |||||||||
| Chymotrypsin C | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Identifiers | |||||||||
| EC number | 3.4.21.2 | ||||||||
| CAS number | 9036-09-3 | ||||||||
| Databases | |||||||||
| IntEnz | IntEnz view | ||||||||
| BRENDA | BRENDA entry | ||||||||
| ExPASy | NiceZyme view | ||||||||
| KEGG | KEGG entry | ||||||||
| MetaCyc | metabolic pathway | ||||||||
| PRIAM | profile | ||||||||
| PDB structures | RCSB PDB PDBe PDBsum | ||||||||
| |||||||||
Structurally, it is the archetypal structure for its superfamily, the PA clan of proteases.
Activation
Chymotrypsin is synthesized in the pancreas by protein biosynthesis as a precursor called chymotrypsinogen that is enzymatically inactive. Trypsin activates chymotrypsinogen by cleaving peptidic bonds in positions Arg15 - Ile16 and produces π-chymotrypsin. In turn, aminic group (-NH3+) of the Ile16 residue interacts with the side chain of Glu194, producing the "oxyanion hole" and the hydrophobic "S1 pocket". Moreover, chymotrypsin induces its own activation by cleaving in positions 14-15, 146-147, and 148-149, producing α-chymotrypsin (which is more active and stable than π-chymotrypsin). The resulting molecule is a three-polypeptide molecule interconnected via disulfide bonds.
Mechanism of action and kinetics
In vivo, chymotrypsin is a proteolytic enzyme (serine protease) acting in the digestive systems of many organisms. It facilitates the cleavage of peptide bonds by a hydrolysis reaction, which despite being thermodynamically favorable, occurs extremely slowly in the absence of a catalyst. The main substrates of chymotrypsin are peptide bonds in which the amino acid N-terminal to the bond is a tryptophan, tyrosine, phenylalanine, or leucine. Like many proteases, chymotrypsin also hydrolyses amide bonds in vitro, a virtue that enabled the use of substrate analogs such as N-acetyl-L-phenylalanine p-nitrophenyl amide for enzyme assays.
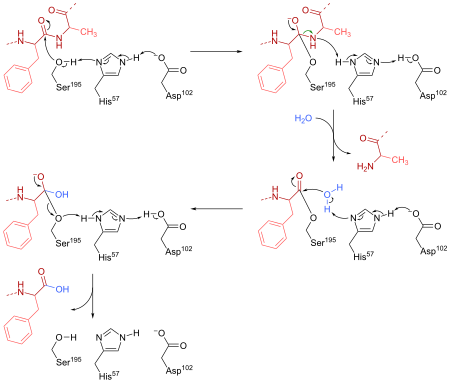
Chymotrypsin cleaves peptide bonds by attacking the unreactive carbonyl group with a powerful nucleophile, the serine 195 residue located in the active site of the enzyme, which briefly becomes covalently bonded to the substrate, forming an enzyme-substrate intermediate. Along with histidine 57 and aspartic acid 102, this serine residue constitutes the catalytic triad of the active site.
These findings rely on inhibition assays and the study of the kinetics of cleavage of the aforementioned substrate, exploiting the fact that the enzyme-substrate intermediate p-nitrophenolate has a yellow colour, enabling measurement of its concentration by measuring light absorbance at 410 nm.
The reaction of chymotrypsin with its substrate was found to take place in two stages, an initial “burst” phase at the beginning of the reaction and a steady-state phase following Michaelis-Menten kinetics. The mode of action of chymotrypsin explains this as hydrolysis takes place in two steps. First, acylation of the substrate to form an acyl-enzyme intermediate, and then deacylation to return the enzyme to its original state. This occurs via the concerted action of the three-amino-acid residues in the catalytic triad.[5] Aspartate hydrogen bonds to the N-δ hydrogen of histidine, increasing the pKa of its ε nitrogen, thus making it able to deprotonate serine. This deprotonation allows the serine side chain to act as a nucleophile and bind to the electron-deficient carbonyl carbon of the protein main chain. Ionization of the carbonyl oxygen is stabilized by formation of two hydrogen bonds to adjacent main chain N-hydrogens. This occurs in the oxyanion hole. This forms a tetrahedral adduct and breakage of the peptide bond. An acyl-enzyme intermediate, bound to the serine, is formed, and the newly formed amino terminus of the cleaved protein can dissociate. In the second reaction step, a water molecule is activated by the basic histidine, and acts as a nucleophile. The oxygen of water attacks the carbonyl carbon of the serine-bound acyl group, resulting in formation of a second tetrahedral adduct, regeneration of the serine -OH group, and release of a proton, as well as the protein fragment with the newly formed carboxyl terminus [5]
Isozymes
|
|
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
See also
- Trypsin
- PA clan of proteases
References
- PDB: 1CHG; Freer ST, Kraut J, Robertus JD, Wright HT, Xuong NH (April 1970). "Chymotrypsinogen: 2.5-angstrom crystal structure, comparison with alpha-chymotrypsin, and implications for zymogen activation". Biochemistry. 9 (9): 1997–2009. doi:10.1021/bi00811a022. PMID 5442169.
- Wilcox PE (1970). "[5] Chymotrypsinogens—chymotrypsins". Chymotrypsinogens — chymotrypsins. Methods in Enzymology. 19. pp. 64–108. doi:10.1016/0076-6879(70)19007-0. ISBN 978-0-12-181881-4.
- Appel W (December 1986). "Chymotrypsin: molecular and catalytic properties". Clin. Biochem. 19 (6): 317–22. doi:10.1016/S0009-9120(86)80002-9. PMID 3555886.
- Berger A, Schechter I (February 1970). "Mapping the active site of papain with the aid of peptide substrates and inhibitors". Philos. Trans. R. Soc. Lond. B Biol. Sci. 257 (813): 249–64. doi:10.1098/rstb.1970.0024. PMID 4399049. S2CID 6877875.
- Petsko, Gregory; Ringe, Dagmar (2009). Protein Structure and Function. Oxford: Oxford University Press. pp. 78–79. ISBN 978-0-19-955684-7.
Further reading
- Stryer L, Berg JM, Tymoczko JL (2002). Biochemistry. San Francisco: W.H. Freeman. ISBN 0-7167-4684-0.
- Grisham CM, Reginald H (2005). Biochemistry. Australia: Thomson Brooks/Cole. ISBN 0-534-49033-6.
External links
- The MEROPS online database for peptidases and their inhibitors: S01.001
- Chymotrypsin at the US National Library of Medicine Medical Subject Headings (MeSH)