Pentapeptide repeat
Pentapeptide repeats are a family of sequence motifs found in multiple tandem copies in protein molecules.[2][3] Pentapeptide repeat proteins are found in all species, but they are found in many copies in cyanobacterial genomes. The repeats were first identified by Black and colleagues in the hglK protein.[4] The later Bateman et al. showed that a large family of related pentapeptide repeat proteins existed.[3] The function of these repeats is uncertain in most proteins. However, in the MfpA protein a DNA gyrase inhibitor it has been suggested that the pentapeptide repeat structure mimics the structure of DNA.[5] The repeats form a regular right handed four sided beta helix structure known as the Rfr-fold.
| Pentapeptide repeat | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
 Structure of the pentapeptide repeat protein HetL.[1] | |||||||||||
| Identifiers | |||||||||||
| Symbol | Pentapeptide | ||||||||||
| Pfam | PF00805 | ||||||||||
| InterPro | IPR001646 | ||||||||||
| |||||||||||
Sequence features

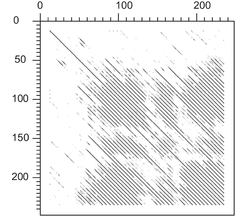
The pentapeptide repeat is a feature seen in protein sequence. It can be approximately described using the 1-letter amino acid code as A(D/N)LXX, where X can be any amino acid . This repeating sequence can be seen in multiple sequence alignments and dot plots of proteins such as HglK. The central position in the pentapeptide repeat is usually a leucine and has been designated as position i. The two previous positions are known as i-1 and i-2. Position i-2 is usually an alanine. The two subsequent positions are denoted i+1 and i+2. The side chains of positions i-2 and i point into the hydrophobic interior of the protein while the side chains of positions i-1, i+1 and i+2 are exposed on the surface of the proteins.
Structure
Pentapeptide repeats were initially predicted from sequence to possess a right handed beta helix with three sides.[3] The first crystal structure of a pentapeptide repeat protein was the MfpA protein solved by Hegde and colleagues. It showed that pentapeptide repeat proteins (PRPs) possessed a four sided beta helix structure.[5] Four repeats make up one turn of a solenoid like structure. The structures of eight different proteins have been solved to date.
| Protein | PDB code | Length | Number of repeats | Reference |
|---|---|---|---|---|
| Mycobacterium tuberculosis MfpA | PDB: 2bm4 | 183 | 30 | [5] |
| Cyanobacterium nostoc HetL | PDB: 3du1 | 237 | 40 | [1] |
| Enterococcus faecalis EfsQnr | PDB: 2w7z | 211 | [6] | |
| Nostoc punctiforme Np275 | PDB: 2J8I | 98 | 17 | [7] |
| Nostoc punctiforme Np276 | PDB: 2J8K | 75 | 12 | [7] |
| Cyanothece sp. Rfr32 | PDB: 2F3L PDB: 2G0Y | 167 | 21 | [8] |
| Cyanothece sp. Rfr23 | PDB: 2O6W | 174 | 23 | [9] |
| Arabidopsis thaliana At2g44920 | PDB: 3N90 | 224 | 25 | [10] |
References
- Ni S, Sheldrick GM, Benning MM, Kennedy MA (January 2009). "The 2 Å resolution crystal structure of HetL, a pentapeptide repeat protein involved in regulation of heterocyst differentiation in the cyanobacterium Nostoc sp. strain PCC 7120". J. Struct. Biol. 165 (1): 47–52. doi:10.1016/j.jsb.2008.09.010. PMID 18952182.
- Vetting MW, Hegde SS, Fajardo JE, et al. (January 2006). "Pentapeptide repeat proteins". Biochemistry. 45 (1): 1–10. doi:10.1021/bi052130w. PMC 2566302. PMID 16388575.
- Bateman A, Murzin AG, Teichmann SA (June 1998). "Structure and distribution of pentapeptide repeats in bacteria". Protein Sci. 7 (6): 1477–80. doi:10.1002/pro.5560070625. PMC 2144021. PMID 9655353.
- Black K, Buikema WJ, Haselkorn R (November 1995). "The hglK gene is required for localization of heterocyst-specific glycolipids in the cyanobacterium Anabaena sp. strain PCC 7120". J. Bacteriol. 177 (22): 6440–8. doi:10.1128/jb.177.22.6440-6448.1995. PMC 177493. PMID 7592418.
- Hegde SS, Vetting MW, Roderick SL, et al. (June 2005). "A fluoroquinolone resistance protein from Mycobacterium tuberculosis that mimics DNA". Science. 308 (5727): 1480–3. doi:10.1126/science.1110699. PMID 15933203.
- Vetting MW, Hegde SS, Blanchard JS (May 2009). "Crystallization of a pentapeptide-repeat protein by reductive cyclic pentylation of free amines with glutaraldehyde". Acta Crystallogr. D. 65 (Pt 5): 462–9. doi:10.1107/S0907444909008324. PMC 2672816. PMID 19390151.
- Vetting MW, Hegde SS, Hazleton KZ, Blanchard JS (April 2007). "Structural characterization of the fusion of two pentapeptide repeat proteins, Np275 and Np276, from Nostoc punctiforme: resurrection of an ancestral protein". Protein Sci. 16 (4): 755–60. doi:10.1110/ps.062637707. PMC 2203339. PMID 17384236.
- Buchko GW, Ni S, Robinson H, Welsh EA, Pakrasi HB, Kennedy MA (November 2006). "Characterization of two potentially universal turn motifs that shape the repeated five-residues fold--crystal structure of a lumenal pentapeptide repeat protein from Cyanothece 51142". Protein Sci. 15 (11): 2579–95. doi:10.1110/ps.062407506. PMC 2242410. PMID 17075135.
- Buchko GW, Robinson H, Pakrasi HB, Kennedy MA (April 2008). "Insights into the structural variation between pentapeptide repeat proteins--crystal structure of Rfr23 from Cyanothece 51142". J. Struct. Biol. 162 (1): 184–92. doi:10.1016/j.jsb.2007.11.008. PMID 18158251.
- Ni S, McGookey ME, Tinch SL, et al. (December 2011). "The 1.7 Å resolution structure of At2g44920, a pentapeptide-repeat protein in the thylakoid lumen of Arabidopsis thaliana". Acta Crystallographica Section F. 67 (Pt 12): 1480–4. doi:10.1107/S1744309111037432. PMC 3232121. PMID 22139148.