MDC1
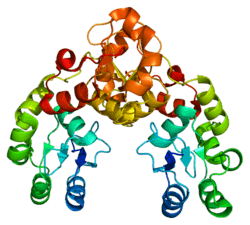
Mediator of DNA damage checkpoint protein 1 is a 2080 amino acid long protein that in humans is encoded by the MDC1 gene[5][6][7] located on the short arm (p) of chromosome 6. MDC1 protein is a regulator of the Intra-S phase and the G2/M cell cycle checkpoints and recruits repair proteins to the site of DNA damage. It is involved in determining cell survival fate in association with tumor suppressor protein p53. This protein also goes by the name Nuclear Factor with BRCT Domain 1 (NFBD1).
Function
Role in DNA damage response
The MDC1 gene encodes the MDC1 nuclear protein which is part of the DNA damage response (DDR) pathway, the mechanism through which eukaryotic cells respond to damaged DNA, specifically DNA double-strand breaks (DSB) that are caused by ionizing radiation or chemical clastogens.[8] The DDR of mammalian cells is made up of kinases, and mediator/adaptors factors.[9] In mammalian cells the DRR is a network of pathways made up of proteins that function as either kinases, or and mediator/adaptors that recruit the kinases to their phosphorylation targets, these factors work together to detect DNA damage, and signal the repair mechanism as well as activating cell cycle checkpoints.[9] The MDC1s role in DDR is to function both as a mediator/adaptor protein mediating a complex of other DDR proteins at the site of DNA damage[9] and repairing DNA damage through its PST domain.[10]
When a cell is exposed to ionizing radiation, its chromatin can be damaged with DSB, triggering the DDR which starts with the MRN complex recruiting ATM kinase to the exposed H2AX histones on the damaged DNA. ATM phosphorylates the C-terminus of the H2AX histone (phosphorylated H2AX histones are commonly noted as γH2AX), and they become an epigenetic flag that highlights the site of DNA damage . The SDT domain of the MDC1 protein is phosphorylated by caseine kinase 2 (CK2) which allows it to bind another MRN complex, the MDC1 protein can sense the DNA damage by binding to the γH2AX flag through its BRCT domain and brings the bound MRN complex to the site of damaged DNA and it facilitates the recruitment and retention of another ATM kinase. The second ATM kinase phosphorylates the TQXF domain on MDC1 which allows it to recruit the E3 ubiquitin ligase RNF8, which will ubiquitinate the histones near the DSB which initiates further ubiquitination of the chromatin around the site of damage by other factors of the DDR. This aggregation of DDR factors and concentration of phosphorylated and ubiquitinated histones is called a DNA damage foci or ionizing radiation-induced foci[9] and the main role of MDC1 is to coordinate the creation of DNA damage foci. This protein is required to activate the intra-S phase and G2/M phase cell cycle checkpoints in response to DNA damage.
Role in apoptosis
MDC1 has anti-apoptotic properties by directly inhibiting the apoptotic activity of the tumor suppressing protein p53. DNA damage can induce apoptosis when the ATM kinase and Chk2 phosphorylate p53 on its Ser-15 and Ser-20 residues which activates p53 and stabilizes it by allowing it to dissociate from the E3 ubiquitin protein ligase MDM2.[11] MDC1 can execute its anti-apoptotic activity by inhibiting p53 in two ways. The MDC1 protein can bind to the n-terminus of p53 through its BRC1 domain which blocks p53 transactivation domain. MDC1 can also inactivate p53 by reducing the phosphorylation levels of p53 Ser-15 residues necessary to p53 apoptotic activity. Studies on lung cancer cell lines (A549 cells) showed an increase in apoptosis in response to genotoxic agents when MDC1 protein levels were reduced with siRNA.[11]
Loss of MDC1 protein
Inhibition or loss of MDC1 protein through studies with siRNA on human cells or knockout studies in mice have shown several defects at both the cellular and organismal level. Mice lacking MDC1 are smaller, have infertile males, are radiosensitive, and are more susceptible to tumors. Knock out MDC1 mice cells and silenced human cells were radiosensitive, failed to initiate Intra-S phase and G2/M checkpoints, failed to produce ionizing radiation-induced foci had poor phosphorylation by the DRR kinases (ATM, CHK1, CHK2), defects in homologous recombination. Human cells with silenced MDC1 also displayed random plasmid integration, reduced apoptosis, and slowed mitosis.[9]
Interactions
MDC1 has been shown to interact with:
MDC1 also binds to mRNA or polyadenylated RNA in the nucleus.[14]
Protein structure
The MDC1 protein contains the following domains listed in order from N-terminal to C-terminal:
- forkhead-associated domain (FHA), N-terminus domain lies between amino acid residues 54 and 105
- SDT (or SDTD) - This domain is located between amino acids 218 and 460.
- TQXF- This domain is located between amino acids 699 and 768.
- PST- This domain lies between amino acid residues 114 and 1662.
- BRCA 1 C-terminus (BRCT) domain and lies between amino acids 1891 and 2082.
- FHA domain
- Unlike the FHA domains on other DRR factors, the FHA domain on MDC1 is not well-characterized. It has been implicated in DSB repair, Intra-S phase and G2/M checkpoints but the specific mechanism is yet to be determined. The FHA domain does have a few putative MDC1-FHA interacting factors such as ATM, CHK2, and RAD51.[9]
- SDT domain
- When the SDT domain is phosphorylated it can bind the MRN complex (composed of MRE11/RAD50/NBS1)[15] and is responsible for keeping the MRN complex associated with the DSB chromatin.[16][17][18] This domain along with NBS1 of the MRN complex are necessary for the activation of intra-S-phase and the G2/M checkpoints, however their role in the molecular mechanism of checkpoint control has not been resolved.[9]
- TQXF domain
- This domain is characterized by four threonine-glutamine then a phenylalanine at the 3+ position.[9] ATM phosphorylates this domain allowing it to bind RNF8 an E3 ubiquitin ligase. This MDC1/RNF8 coupling then facilitates the recruitment of other DDR factors such as RNF168, 53BP1, and BRCA1.[9] TQXF is important for proper passage through the G2/M checkpoint, however the molecular mechanism through which MDC1 and RNF8 regulate the G2/M checkpoint has not yet been resolved.
- PST domain
- The PST domain is composed of repeats of a proline-serine-threonine motif. This domain plays a role in DNA repair by both homologous recombination and by Non-homologous end joining, however the mechanism through which it facilitates repair of damaged DNA is not yet known.[10]
- BRCT domain
- The BRCT domain on MDC1 directly binds to the γH2AX of damaged chromatin. The BRCT domain creates an α/β fold which extends from the C-terminus of MDC1 through a linker region. It preferentially binds to phosphorylated Ser residues followed by Glu,Tyr, motif on γH2AX.[19] This domain also binds to anaphase-promoting complex (APC/C) which is an E3 ubiquitin ligase that degrades cyclins.[20] The BRCT domain is involved in the regulation of the decatenation checkpoint at the end of replication by binding Topo IIα this arrest the cell in the G2 cycle until the sister chromatin have completely separated.[21] The BRCT domain also interacts the tumor suppressor p53 and inhibits p53 by blocking its transactivation domain, as well as aiding in MDM2 inactivation of p53.[11]
Regulation
MDC1 is indirectly down regulated by the oncogene AKT1. AKT1 activates expression of the microRNA-22 (miR-22) which targets the 3’ end of MDC1 mRNA inhibiting translation. Aberrant overexpression of AKT1, which is observed in several cancers including breast, lung and prostate, results in reduced production of MDC1 and subsequently a destabilization of the genome and increased tumorigenicity.[22]
Role in cancer
MDC1 is a putative tumor suppressor. Knockout studies in mice have shown an increase in tumor development when MDC1 is lost. Reduction in MDC1 protein levels has been observed in a large number of breast and lung carcinomas.[23][24] Several studies on various human cancer cell lines including the A549 cell human lung carcinoma line,[11] multiple esophageal cancer cell lines (TE11, YES2 ,YES5),[25] and cervical cancer cell lines (HeLa, SiHa, and CaSki)[26] showed increased sensitivity to anti-cancer drugs (adriamycin and cisplatin), when endogenous MDC1 protein levels were knockdown with siRNA. Because of MDC1s involvement in several pathways that are often misappropriated by cancer cells including the cell cycle checkpoints, DDR, and p53 tumor suppression, cancer treatments that target MDC1 have the potential to be potent radiosensitizer and chemosensitizer.
References
- ENSG00000237095, ENSG00000137337, ENSG00000206481, ENSG00000234012, ENSG00000231135, ENSG00000228575, ENSG00000225589 GRCh38: Ensembl release 89: ENSG00000224587, ENSG00000237095, ENSG00000137337, ENSG00000206481, ENSG00000234012, ENSG00000231135, ENSG00000228575, ENSG00000225589 - Ensembl, May 2017
- GRCm38: Ensembl release 89: ENSMUSG00000061607 - Ensembl, May 2017
- "Human PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- "Mouse PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- Ozaki T, Nagase T, Ichimiya S, Seki N, Ohiri M, Nomura N, Takada N, Sakiyama S, Weber BL, Nakagawara A (Sep 2000). "NFBD1/KIAA0170 is a novel nuclear transcriptional transactivator with BRCT domain". DNA Cell Biol. 19 (8): 475–485. doi:10.1089/10445490050128403. PMID 10975465.
- Stewart GS, Wang B, Bignell CR, Taylor AM, Elledge SJ (Feb 2003). "MDC1 is a mediator of the mammalian DNA damage checkpoint". Nature. 421 (6926): 961–966. doi:10.1038/nature01446. PMID 12607005.
- "Entrez Gene: MDC1 mediator of DNA damage checkpoint 1".
- Zhou BB, Elledge SJ (Nov 2000). "The DNA damage response: putting checkpoints in perspective". Nature. 408 (6811): 433–439. doi:10.1038/35044005. PMID 11100718.
- Jungmichel S, Stucki M (Aug 2010). "MDC1: The art of keeping things in focus" (PDF). Chromosoma. 119 (4): 337–349. doi:10.1007/s00412-010-0266-9. PMID 20224865.
- Lou Z, Chen BP, Asaithamby A, Minter-Dykhouse K, Chen DJ, Chen J (Nov 2004). "MDC1 regulates DNA-PK autophosphorylation in response to DNA damage". J Biol Chem. 279 (45): 46359–62. doi:10.1074/jbc.c400375200. PMID 15377652.
- Nakanishi M, Ozaki T, Yamamoto H, Hanamoto T, Kikuchi H, Furuya K, Asaka M, Delia D, Nakagawara A (Aug 2007). "NFBD1/MDC1 associates with p53 and regulates its function at the crossroad between cell survival and death in response to DNA damage". J Biol Chem. 282 (31): 22993–3004. doi:10.1074/jbc.m611412200. PMID 17535811.
- Lou Z, Minter-Dykhouse K, Wu X, Chen J (Feb 2003). "MDC1 is coupled to activated CHK2 in mammalian DNA damage response pathways". Nature. 421 (6926): 957–61. doi:10.1038/nature01447. PMID 12607004.
- Xu X, Stern DF (Oct 2003). "NFBD1/MDC1 regulates ionizing radiation-induced focus formation by DNA checkpoint signaling and repair factors". FASEB J. 17 (13): 1842–8. doi:10.1096/fj.03-0310com. PMID 14519663.
- Conrad T, Albrecht AS, de Melo Costa VR, Sauer S, Meierhofer D, Ørom UA (2016-01-01). "Serial interactome capture of the human cell nucleus". Nature Communications. 7: 11212. doi:10.1038/ncomms11212. PMC 4822031. PMID 27040163.
- de Jager M, van Noort J, van Gent DC, Dekker C, Kanaar R, Wyman C (Nov 2001). "Human Rad50/Mre11 is a flexible complex that can tether DNA ends". Mol Cell. 8 (5): 1129–1135. doi:10.1016/s1097-2765(01)00381-1. PMID 11741547.
- Bekker-Jensen S, Lukas C, Kitagawa R, Melander F, Kastan MB, Bartek J, Lukas J (Apr 2006). "Spatial organization of the mammalian genome surveillance machinery in response to DNA strand breaks". J Cell Biol. 173 (2): 195–206. doi:10.1083/jcb.200510130. PMC 2063811. PMID 16618811.
- Goldberg M, Stucki M, Falck J, D'amours D, Rahman D, Pappin D, Bartek J, Jackson S (Feb 2003). "MDC1 is required for the intra-S- phase DNA damage checkpoint". Nature. 421 (6926): 952–956. doi:10.1038/nature01445. PMID 12607003.
- Lukas C, Melander F, Stucki M, Falck J, Bekker-Jensen S, Goldberg M, Lerenthal Y, Jackson S, Bartek J, Lukas J (Jul 2004). "Mdc1 couples DNA double-strand break recognition by Nbs1 with its H2AX- dependent chromatin retention". EMBO J. 23 (13): 2674–2683. doi:10.1038/sj.emboj.7600269. PMC 449779. PMID 15201865.
- Lou Z, Minter-Dykhouse K, Franco S, Gostissa M, Rivera MA, Celeste A, Manis JP, van Deursen J, Nussenzweig A, Paull TT, Alt FW, Chen J (Jan 2006). "MDC1 maintains genomic stability by participating in the amplification of ATM-dependent DNA damage signals". Mol Cell. 21 (2): 187–200. doi:10.1016/j.molcel.2005.11.025. PMID 16427009.
- Coster G, Hayouka Z, Argaman L, Strauss C, Friedler A, Brandeis M, Goldberg M (Sep 2007). "The DNA damage response mediator MDC1 directly interacts with the anaphase-promoting complex/cyclosome". J Biol Chem. 282 (44): 32053–32064. doi:10.1074/jbc.m705890200. PMID 17827148.
- Luo K, Yuan J, Chen J, Lou Z (Feb 2009). "Topoisomerase IIalpha controls the decatenation checkpoint". Nat Cell Biol. 11 (2): 204–210. doi:10.1038/ncb1828. PMC 2712943. PMID 19098900.
- Lee JH, Park SJ, Jeong SY, Kim MJ, Jun S, Lee HS, Chang IY, Lim SC, Yoon SP, Yong J, and You HJ (Apr 2015). "MicroRNA-22 Suppresses DNA Repair and Promotes Genomic Instability through Targeting of MDC1". Cancer Research. 75 (7): 1298–1310. doi:10.1158/0008-5472.CAN-14-2783. PMID 25627978.
- Minter-Dykhouse K, Ward I, Huen SY, Chen J, Lou Z (Jun 2008). "Distinct versus overlapping functions of MDC1 and 53BP1 in DNA damage response and tumorigenesis". J Cell Biol. 181 (5): 727–735. doi:10.1083/jcb.200801083. PMC 2396806. PMID 18504301.
- Bartkova J, Hořejsí Z, Sehested M, Nesland JM, Rajpert-De Meyts E, Skakkebæk NE, Stucki M, Jackson S, Lukas J, Bartek J (Jun 2007). "DNA damage response mediators MDC1 and 53BP1: constitu- tive activation and aberrant loss in breast and lung cancer, but not in testicular germ cell tumours". Oncogene. 26 (53): 7414–7422. doi:10.1038/sj.onc.1210553. PMID 17546051.
- Yang M, Bu Y, Wang C, Liu G, Song F (Sep 2010). "Growth inhibition, morphology change, and cell cycle alterations in NFBD1-depleted human esophageal cancer cells". Mol Cell Biochem. 342 (1–2): 1–6. doi:10.1007/s11010-010-0460-3. PMID 20364298.
- Yuan C, Bu Y, Wang C, Yi F, Yang Z, Huang X, Cheng L, Liu G, Wang Y, Song F (Jan 2010). "NFBD1/MDC1 is a protein of oncogenic potential in human cervical cancer". Mol Cell Biochem. 359 (1–2): 333–46. doi:10.1007/s11010-011-1027-7. PMID 21853275.
Further reading
- Stucki M, Jackson SP (2005). "MDC1/NFBD1: a key regulator of the DNA damage response in higher eukaryotes". DNA Repair (Amst.). 3 (8–9): 953–957. doi:10.1016/j.dnarep.2004.03.007. PMID 15279781.
- Nagase T, Seki N, Ishikawa K, Tanaka A, Nomura N (1996). "Prediction of the coding sequences of unidentified human genes. V. The coding sequences of 40 new genes (KIAA0161-KIAA0200) deduced by analysis of cDNA clones from human cell line KG-1". DNA Res. 3 (1): 17–24. doi:10.1093/dnares/3.1.17. PMID 8724849.
- Shang YL, Bodero AJ, Chen PL (2003). "NFBD1, a novel nuclear protein with signature motifs of FHA and BRCT, and an internal 41-amino acid repeat sequence, is an early participant in DNA damage response". J. Biol. Chem. 278 (8): 6323–6329. doi:10.1074/jbc.M210749200. PMID 12475977.
- Xu X, Stern DF (2003). "NFBD1/KIAA0170 is a chromatin-associated protein involved in DNA damage signaling pathways". J. Biol. Chem. 278 (10): 8795–8803. doi:10.1074/jbc.M211392200. PMID 12499369.
- Peng A, Chen PL (2003). "NFBD1, like 53BP1, is an early and redundant transducer mediating Chk2 phosphorylation in response to DNA damage". J. Biol. Chem. 278 (11): 8873–8876. doi:10.1074/jbc.C300001200. PMID 12551934.
- Goldberg M, Stucki M, Falck J, D'Amours D, Rahman D, Pappin D, Bartek J, Jackson SP (2003). "MDC1 is required for the intra-S-phase DNA damage checkpoint". Nature. 421 (6926): 952–956. doi:10.1038/nature01445. PMID 12607003.
- Lou Z, Minter-Dykhouse K, Wu X, Chen J (2003). "MDC1 is coupled to activated CHK2 in mammalian DNA damage response pathways". Nature. 421 (6926): 957–961. doi:10.1038/nature01447. PMID 12607004.
- Lou Z, Chini CC, Minter-Dykhouse K, Chen J (2003). "Mediator of DNA damage checkpoint protein 1 regulates BRCA1 localization and phosphorylation in DNA damage checkpoint control". J. Biol. Chem. 278 (16): 13599–13602. doi:10.1074/jbc.C300060200. PMID 12611903.
- Xu X, Stern DF (2003). "NFBD1/MDC1 regulates ionizing radiation-induced focus formation by DNA checkpoint signaling and repair factors". FASEB J. 17 (13): 1842–1848. doi:10.1096/fj.03-0310com. PMID 14519663.
- Rodriguez M, Yu X, Chen J, Songyang Z (2004). "Phosphopeptide binding specificities of BRCA1 COOH-terminal (BRCT) domains". J. Biol. Chem. 278 (52): 52914–52918. doi:10.1074/jbc.C300407200. PMID 14578343.
- Mochan TA, Venere M, DiTullio RA, Halazonetis TD (2004). "53BP1 and NFBD1/MDC1-Nbs1 function in parallel interacting pathways activating ataxia-telangiectasia mutated (ATM) in response to DNA damage". Cancer Res. 63 (24): 8586–91. PMID 14695167.
- Lukas C, Melander F, Stucki M, Falck J, Bekker-Jensen S, Goldberg M, Lerenthal Y, Jackson SP, Bartek J, Lukas J (2005). "Mdc1 couples DNA double-strand break recognition by Nbs1 with its H2AX-dependent chromatin retention". EMBO J. 23 (13): 2674–2683. doi:10.1038/sj.emboj.7600269. PMC 449779. PMID 15201865.
- Beausoleil SA, Jedrychowski M, Schwartz D, Elias JE, Villén J, Li J, Cohn MA, Cantley LC, Gygi SP (2004). "Large-scale characterization of HeLa cell nuclear phosphoproteins". Proc. Natl. Acad. Sci. U.S.A. 101 (33): 12130–12135. doi:10.1073/pnas.0404720101. PMC 514446. PMID 15302935.
- Lou Z, Chen BP, Asaithamby A, Minter-Dykhouse K, Chen DJ, Chen J (2004). "MDC1 regulates DNA-PK autophosphorylation in response to DNA damage". J. Biol. Chem. 279 (45): 46359–46362. doi:10.1074/jbc.C400375200. PMID 15377652.
- Polci R, Peng A, Chen PL, Riley DJ, Chen Y (2005). "NIMA-related protein kinase 1 is involved early in the ionizing radiation-induced DNA damage response". Cancer Res. 64 (24): 8800–8803. doi:10.1158/0008-5472.CAN-04-2243. PMID 15604234.