H2AFX
H2A histone family member X (usually abbreviated as H2AX) is a type of histone protein from the H2A family encoded by the H2AFX gene. An important phosphorylated form is γH2AX (140S), which forms when double-strand breaks appear.
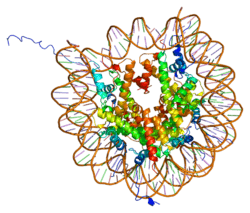
In humans and other eukaryotes, the DNA is wrapped around histone octamers, consisting of core histones H2A, H2B, H3 and H4, to form chromatin. H2AX contributes to nucleosome-formation, chromatin-remodeling and DNA repair, and is also used in vitro as an assay for double-strand breaks in dsDNA.
Formation of γH2AX
H2AX becomes phosphorylated on serine 139, then called γH2AX, as a reaction on DNA double-strand breaks (DSB). The kinases of the PI3-family (Ataxia telangiectasia mutated, ATR and DNA-PKcs) are responsible for this phosphorylation, especially ATM. The modification can happen accidentally during replication fork collapse or in the response to ionizing radiation but also during controlled physiological processes such as V(D)J recombination. γH2AX is a sensitive target for looking at DSBs in cells. The presence of γH2AX by itself, however, is not the evidence of the DSBs.[5][6] The role of the phosphorylated form of the histone in DNA repair is under discussion but it is known that because of the modification the DNA becomes less condensed, potentially allowing space for the recruitment of proteins necessary during repair of DSBs. Mutagenesis experiments have shown that the modification is necessary for the proper formation of ionizing radiation induced foci in response to double strand breaks, but is not required for the recruitment of proteins to the site of DSBs.
Function
DNA damage response
The histone variant H2AX constitutes about 2-25% of the H2A histones in mammalian chromatin.[7] When a double-strand break occurs in DNA, a sequence of events occurs in which H2AX is altered.
Very early after a double-strand break, a specific protein that interacts with and affects the architecture of chromatin is phosphorylated and then released from the chromatin. This protein, heterochromatin protein 1 (HP1)-beta (CBX1), is bound to histone H3 methylated on lysine 9 (H3K9me). Half-maximum release of HP1-beta from damaged DNA occurs within one second.[8] A dynamic alteration in chromatin structure is triggered by HP1-beta release. This alteration in chromatin structure promotes H2AX phosphorylation by ATM, ATR and DNA-PK,[9] allowing formation of γH2AX (H2AX phosphorylated on serine 139). γH2AX can be detected as soon as 20 seconds after irradiation of cells (with DNA double-strand break formation), and half maximum accumulation of γH2AX occurs in one minute.[7] Chromatin with phosphorylated γH2AX extends to about a million base pairs on each side of a DNA double-strand break.[7]
MDC1 (mediator of DNA damage checkpoint protein 1) then binds to γH2AX and the γH2AX/MDC1 complex then orchestrates further interactions in double-strand break repair.[10] The ubiquitin ligases RNF8 and RNF168 bind to the γH2AX/MDC1 complex, ubiquitylating other chromatin components. This allows the recruitment of BRCA1 and 53BP1 to the long, modified γH2AX/MDC1 chromatin.[10] Other proteins that stably assemble on the extensive γH2AX-modified chromatin are the MRN complex (a protein complex consisting of Mre11, Rad50 and Nbs1), RAD51 and the ATM kinase.[11][12] Further DNA repair components, such as RAD52 and RAD54, rapidly and reversibly interact with the core components stably associated with γH2AX-modified chromatin.[12] The constitutive level of γH2AX expression in live cells, untreated by exogenous agents, likely represents DNA damage by endogenous oxidants generated during cellular respiration.[13]
In chromatin remodeling
The packaging of eukaryotic DNA into chromatin presents a barrier to all DNA-based processes that require recruitment of enzymes to their sites of action. To allow DNA repair, the chromatin must be remodeled.
γH2AX, the phosphorylated form of H2AX, is involved in the steps leading to chromatin decondensation after DNA double-strand breaks. γH2AX does not, itself, cause chromatin decondensation, but within 30 seconds of ionizing radiation, RNF8 protein can be detected in association with γH2AX.[14] RNF8 mediates extensive chromatin decondensation, through its subsequent interaction with CHD4,[15] a component of the nucleosome remodeling and deacetylase complex NuRD.
γH2AX as an assay for double-strand breaks
An assay for γH2AX generally reflects the presence of double-strand breaks in DNA, though the assay may indicate other minor phenomena as well.[16] On the one hand, overwhelming evidence supports a strong, quantitative correlation between γH2AX foci formation and DNA double-strand break induction following ionizing radiation exposure, based on absolute yields and distributions induced per unit dose.[16] On the other hand, not only the formation of distinct γH2AX foci but also the induction of pan-nuclear γH2AX signals have been reported as a cellular reaction to various stressors other than ionizing radiation.[17] The γH2AX signal is always stronger at DNA double-strand breaks than in undamaged chromatin.[17] γH2AX in undamaged chromatin is thought to possibly be generated via direct phosphorylation of H2AX by activated kinases, most likely diffusing from DNA damage sites.
Interactions
H2AX has been shown to interact with:
References
- GRCh38: Ensembl release 89: ENSG00000188486 - Ensembl, May 2017
- GRCm38: Ensembl release 89: ENSMUSG00000049932 - Ensembl, May 2017
- "Human PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- "Mouse PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- Cleaver JE, Feeney L, Revet I (2011). "Phosphorylated H2Ax is not an unambiguous marker for DNA double strand breaks". Cell Cycle. 10 (19): 3223–4. doi:10.4161/cc.10.19.17448. PMID 21921674.
- Rybak P, Hoang A, Bujnowicz L, Bernas T, Berniak K, Zarębski M, Darzynkiewicz Z, Dobrucki J (2016). "Low level phosphorylation of histone H2AX on serine 139 (γH2AX) is not associated with DNA double-strand breaks". Oncotarget. 7 (31): 49574–49587. doi:10.18632/oncotarget.10411. PMC 5226530. PMID 27391338.
- Rogakou EP, Pilch DR, Orr AH, Ivanova VS, Bonner WM (1998). "DNA double-stranded breaks induce histone H2AX phosphorylation on serine 139". J. Biol. Chem. 273 (10): 5858–68. doi:10.1074/jbc.273.10.5858. PMID 9488723.
- Ayoub N, Jeyasekharan AD, Bernal JA, Venkitaraman AR (2008). "HP1-beta mobilization promotes chromatin changes that initiate the DNA damage response". Nature. 453 (7195): 682–6. Bibcode:2008Natur.453..682A. doi:10.1038/nature06875. PMID 18438399.
- Furuta T, Takemura H, Liao ZY, Aune GJ, Redon C, Sedelnikova OA, Pilch DR, Rogakou EP, Celeste A, Chen HT, Nussenzweig A, Aladjem MI, Bonner WM, Pommier Y (2003). "Phosphorylation of histone H2AX and activation of Mre11, Rad50, and Nbs1 in response to replication-dependent DNA double-strand breaks induced by mammalian DNA topoisomerase I cleavage complexes". J. Biol. Chem. 278 (22): 20303–12. doi:10.1074/jbc.M300198200. PMID 12660252.
- Scully R, Xie A (2013). "Double strand break repair functions of histone H2AX". Mutat. Res. 750 (1–2): 5–14. doi:10.1016/j.mrfmmm.2013.07.007. PMC 3818383. PMID 23916969.
- Bekker-Jensen S, Lukas C, Kitagawa R, Melander F, Kastan MB, Bartek J, Lukas J (2006). "Spatial organization of the mammalian genome surveillance machinery in response to DNA strand breaks". J. Cell Biol. 173 (2): 195–206. doi:10.1083/jcb.200510130. PMC 2063811. PMID 16618811.
- Essers J, Houtsmuller AB, van Veelen L, Paulusma C, Nigg AL, Pastink A, Vermeulen W, Hoeijmakers JH, Kanaar R (2002). "Nuclear dynamics of RAD52 group homologous recombination proteins in response to DNA damage". EMBO J. 21 (8): 2030–7. doi:10.1093/emboj/21.8.2030. PMC 125370. PMID 11953322.
- Tanaka T, Halicka HD, Huang X, Traganos F, Darzynkiewicz Z (2006). "Constitutive histone H2AX phosphorylation and ATM activation, the reporters of DNA damage by endogenous oxidants". Cell Cycle. 5 (17): 1940–5. doi:10.4161/cc.5.17.3191. PMC 3488278. PMID 16940754.
- Mailand N, Bekker-Jensen S, Faustrup H, Melander F, Bartek J, Lukas C, Lukas J (2007). "RNF8 ubiquitylates histones at DNA double-strand breaks and promotes assembly of repair proteins". Cell. 131 (5): 887–900. doi:10.1016/j.cell.2007.09.040. PMID 18001824.
- Luijsterburg MS, Acs K, Ackermann L, Wiegant WW, Bekker-Jensen S, Larsen DH, Khanna KK, van Attikum H, Mailand N, Dantuma NP (2012). "A new non-catalytic role for ubiquitin ligase RNF8 in unfolding higher-order chromatin structure". EMBO J. 31 (11): 2511–27. doi:10.1038/emboj.2012.104. PMC 3365417. PMID 22531782.
- Rothkamm K, Barnard S, Moquet J, Ellender M, Rana Z, Burdak-Rothkamm S (2015). "DNA damage foci: Meaning and significance". Environ. Mol. Mutagen. 56 (6): 491–504. doi:10.1002/em.21944. PMID 25773265.
- Meyer B, Voss KO, Tobias F, Jakob B, Durante M, Taucher-Scholz G (2013). "Clustered DNA damage induces pan-nuclear H2AX phosphorylation mediated by ATM and DNA-PK". Nucleic Acids Res. 41 (12): 6109–18. doi:10.1093/nar/gkt304. PMC 3695524. PMID 23620287.
- Mallery DL, Vandenberg CJ, Hiom K (Dec 2002). "Activation of the E3 ligase function of the BRCA1/BARD1 complex by polyubiquitin chains". The EMBO Journal. 21 (24): 6755–62. doi:10.1093/emboj/cdf691. PMC 139111. PMID 12485996.
- Chen A, Kleiman FE, Manley JL, Ouchi T, Pan ZQ (Jun 2002). "Autoubiquitination of the BRCA1*BARD1 RING ubiquitin ligase". The Journal of Biological Chemistry. 277 (24): 22085–92. doi:10.1074/jbc.M201252200. PMID 11927591.
- Paull TT, Rogakou EP, Yamazaki V, Kirchgessner CU, Gellert M, Bonner WM (2000). "A critical role for histone H2AX in recruitment of repair factors to nuclear foci after DNA damage". Current Biology. 10 (15): 886–95. doi:10.1016/s0960-9822(00)00610-2. PMID 10959836.
- Sengupta S, Robles AI, Linke SP, Sinogeeva NI, Zhang R, Pedeux R, Ward IM, Celeste A, Nussenzweig A, Chen J, Halazonetis TD, Harris CC (Sep 2004). "Functional interaction between BLM helicase and 53BP1 in a Chk1-mediated pathway during S-phase arrest". The Journal of Cell Biology. 166 (6): 801–13. doi:10.1083/jcb.200405128. PMC 2172115. PMID 15364958.
- Stewart GS, Wang B, Bignell CR, Taylor AM, Elledge SJ (Feb 2003). "MDC1 is a mediator of the mammalian DNA damage checkpoint". Nature. 421 (6926): 961–6. Bibcode:2003Natur.421..961S. doi:10.1038/nature01446. PMID 12607005.
- Xu X, Stern DF (Oct 2003). "NFBD1/MDC1 regulates ionizing radiation-induced focus formation by DNA checkpoint signaling and repair factors". FASEB Journal. 17 (13): 1842–8. doi:10.1096/fj.03-0310com. PMID 14519663.
- Kobayashi J, Tauchi H, Sakamoto S, Nakamura A, Morishima K, Matsuura S, Kobayashi T, Tamai K, Tanimoto K, Komatsu K (Oct 2002). "NBS1 localizes to γH2AX foci through interaction with the FHA/BRCT domain". Current Biology. 12 (21): 1846–51. doi:10.1016/s0960-9822(02)01259-9. PMID 12419185.
- Fernandez-Capetillo O, Chen HT, Celeste A, Ward I, Romanienko PJ, Morales JC, Naka K, Xia Z, Camerini-Otero RD, Motoyama N, Carpenter PB, Bonner WM, Chen J, Nussenzweig A (Dec 2002). "DNA damage-induced G2-M checkpoint activation by histone H2AX and 53BP1". Nature Cell Biology. 4 (12): 993–7. doi:10.1038/ncb884. PMID 12447390.
- Ward IM, Minn K, Jorda KG, Chen J (May 2003). "Accumulation of checkpoint protein 53BP1 at DNA breaks involves its binding to phosphorylated histone H2AX". The Journal of Biological Chemistry. 278 (22): 19579–82. doi:10.1074/jbc.C300117200. PMID 12697768.
Further reading
- Redon C, Pilch D, Rogakou E, Sedelnikova O, Newrock K, Bonner W (Apr 2002). "Histone H2A variants H2AX and H2AZ". Current Opinion in Genetics & Development. 12 (2): 162–9. doi:10.1016/S0959-437X(02)00282-4. PMID 11893489.
- Fernandez-Capetillo O, Lee A, Nussenzweig M, Nussenzweig A (2005). "H2AX: the histone guardian of the genome". DNA Repair. 3 (8–9): 959–67. doi:10.1016/j.dnarep.2004.03.024. PMID 15279782.
- Mannironi C, Bonner WM, Hatch CL (Nov 1989). "H2A.X. a histone isoprotein with a conserved C-terminal sequence, is encoded by a novel mRNA with both DNA replication type and polyA 3' processing signals". Nucleic Acids Research. 17 (22): 9113–26. doi:10.1093/nar/17.22.9113. PMC 335118. PMID 2587254.
- Banerjee S, Smallwood A, Hultén M (Feb 1995). "ATP-dependent reorganization of human sperm nuclear chromatin". Journal of Cell Science. 108 (2): 755–65. PMID 7769017.
- Ivanova VS, Hatch CL, Bonner WM (Sep 1994). "Characterization of the human histone H2A.X gene. Comparison of its promoter with other H2A gene promoters". The Journal of Biological Chemistry. 269 (39): 24189–94. PMID 7929075.
- Ivanova VS, Zimonjic D, Popescu N, Bonner WM (Sep 1994). "Chromosomal localization of the human histone H2A.X gene to 11q23.2-q23.3 by fluorescence in situ hybridization". Human Genetics. 94 (3): 303–6. doi:10.1007/BF00208289. PMID 8076949.
- Rogakou EP, Pilch DR, Orr AH, Ivanova VS, Bonner WM (Mar 1998). "DNA double-stranded breaks induce histone H2AX phosphorylation on serine 139". The Journal of Biological Chemistry. 273 (10): 5858–68. doi:10.1074/jbc.273.10.5858. PMID 9488723.
- El Kharroubi A, Piras G, Zensen R, Martin MA (May 1998). "Transcriptional activation of the integrated chromatin-associated human immunodeficiency virus type 1 promoter". Molecular and Cellular Biology. 18 (5): 2535–44. doi:10.1128/mcb.18.5.2535. PMC 110633. PMID 9566873.
- Rogakou EP, Boon C, Redon C, Bonner WM (Sep 1999). "Megabase chromatin domains involved in DNA double-strand breaks in vivo". The Journal of Cell Biology. 146 (5): 905–16. doi:10.1083/jcb.146.5.905. PMC 2169482. PMID 10477747.
- Rogakou EP, Nieves-Neira W, Boon C, Pommier Y, Bonner WM (Mar 2000). "Initiation of DNA fragmentation during apoptosis induces phosphorylation of H2AX histone at serine 139". The Journal of Biological Chemistry. 275 (13): 9390–5. doi:10.1074/jbc.275.13.9390. PMID 10734083.
- Paull TT, Rogakou EP, Yamazaki V, Kirchgessner CU, Gellert M, Bonner WM (2001). "A critical role for histone H2AX in recruitment of repair factors to nuclear foci after DNA damage". Current Biology. 10 (15): 886–95. doi:10.1016/S0960-9822(00)00610-2. PMID 10959836.
- Deng L, de la Fuente C, Fu P, Wang L, Donnelly R, Wade JD, Lambert P, Li H, Lee CG, Kashanchi F (Nov 2000). "Acetylation of HIV-1 Tat by CBP/P300 increases transcription of integrated HIV-1 genome and enhances binding to core histones". Virology. 277 (2): 278–95. doi:10.1006/viro.2000.0593. PMID 11080476.
- Chen HT, Bhandoola A, Difilippantonio MJ, Zhu J, Brown MJ, Tai X, Rogakou EP, Brotz TM, Bonner WM, Ried T, Nussenzweig A (Dec 2000). "Response to RAG-mediated VDJ cleavage by NBS1 and γH2AX". Science. 290 (5498): 1962–5. Bibcode:2000Sci...290.1962C. doi:10.1126/science.290.5498.1962. PMC 4721589. PMID 11110662.
- Chadwick BP, Willard HF (May 2001). "Histone H2A variants and the inactive X chromosome: identification of a second macroH2A variant". Human Molecular Genetics. 10 (10): 1101–13. doi:10.1093/hmg/10.10.1101. PMID 11331621.
- Burma S, Chen BP, Murphy M, Kurimasa A, Chen DJ (Nov 2001). "ATM phosphorylates histone H2AX in response to DNA double-strand breaks". The Journal of Biological Chemistry. 276 (45): 42462–7. doi:10.1074/jbc.C100466200. PMID 11571274.
- Ward IM, Chen J (Dec 2001). "Histone H2AX is phosphorylated in an ATR-dependent manner in response to replicational stress". The Journal of Biological Chemistry. 276 (51): 47759–62. doi:10.1074/jbc.C100569200. PMID 11673449.
- Deng L, Wang D, de la Fuente C, Wang L, Li H, Lee CG, Donnelly R, Wade JD, Lambert P, Kashanchi F (Oct 2001). "Enhancement of the p300 HAT activity by HIV-1 Tat on chromatin DNA". Virology. 289 (2): 312–26. doi:10.1006/viro.2001.1129. PMID 11689053.
- Chen A, Kleiman FE, Manley JL, Ouchi T, Pan ZQ (Jun 2002). "Autoubiquitination of the BRCA1*BARD1 RING ubiquitin ligase". The Journal of Biological Chemistry. 277 (24): 22085–92. doi:10.1074/jbc.M201252200. PMID 11927591.
- Zhu H, Hunter TC, Pan S, Yau PM, Bradbury EM, Chen X (Apr 2002). "Residue-specific mass signatures for the efficient detection of protein modifications by mass spectrometry". Analytical Chemistry. 74 (7): 1687–94. doi:10.1021/ac010853p. PMID 12033261.