5,10-Methenyltetrahydromethanopterin hydrogenase
The 5,10-methenyltetrahydromethanopterin hydrogenase (or Hmd), the so-called iron-sulfur cluster-free hydrogenase, is an enzyme found in methanogenic archea such as Methanothermobacter marburgensis. It was discovered and first characterized by the Thauer group at the Max Planck Institute in Marburg. Hydrogenases are enzymes that either reduce protons or oxidize molecular dihydrogen.[1]
| 5,10-methenyltetrahydromethanopterin hydrogenase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Identifiers | |||||||||
| EC number | 1.12.98.2 | ||||||||
| CAS number | 100357-01-5 | ||||||||
| Databases | |||||||||
| IntEnz | IntEnz view | ||||||||
| BRENDA | BRENDA entry | ||||||||
| ExPASy | NiceZyme view | ||||||||
| KEGG | KEGG entry | ||||||||
| MetaCyc | metabolic pathway | ||||||||
| PRIAM | profile | ||||||||
| PDB structures | RCSB PDB PDBe PDBsum | ||||||||
| Gene Ontology | AmiGO / QuickGO | ||||||||
| |||||||||
| H2-forming N5,N10-methylene-tetrahydromethanopterin dehydrogenase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
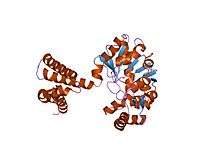 the crystal structure of the apoenzyme of the iron-sulfur-cluster-free hydrogenase (hmd) | |||||||||
| Identifiers | |||||||||
| Symbol | HMD | ||||||||
| Pfam | PF03201 | ||||||||
| InterPro | IPR004889 | ||||||||
| |||||||||
Enzyme function
Methanogens rely on such enzymes to catalyze the reduction of CO2, to methane. One step in methanogenesis entails conversion of a methenyl group (formic acid oxidation state) to a methylene group (formaldehyde oxidation state).
Among the hydrogenase family of enzymes, Hmd is unique in that it does not directly reduce CO2 to CH4. The natural substrate of the enzyme is the organic compound methenyltetrahydromethanopterin.[2] The organic compound includes a methenyl group bound to two tertiary amides. The methenyl group originated as CO2 before being incorporated into the substrate, which is catalytically reduced by H2 to methylenetetrahydromethanopterin as shown.[3] Eventually the methylene group is further reduced and released as a molecule of methane.
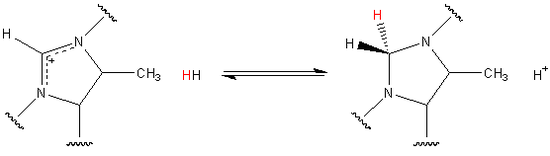
The hydride transfer has also been shown to be stereospecific. Given that the substrate is planar the hydride originating from H2 is always added to the pro-R face. In the reverse reaction stereospecificity is maintained and the highlighted hydride is removed.[1]
Chemical and physical properties
Hmd Holoenzyme
The Hmd holoenzyme includes the protein homodimer as well as its associated iron-containing co-factor. Several species of methanogens have been characterized that express enzymes in the Hmd hydrogenase family. Between species the enzyme is found with differing numbers of sub-units and some minor amino acid sequence variations. The monomer is approximately 45,000 Da in mass, although this value varies from species to species.[2]
The enzymatic activity of the enzyme is lost upon exposure to sunlight or UV. Photolysis causes the release of an iron atom and two molecules of carbon monoxide. In the holoenzyme the Fe and CO molecules are found associated with a 542 Da cofactor.[4]
Hmd iron-containing co-factor
The iron-containing co-factor is found tightly associated with the protein. It can be released upon denaturation with 2-mercaptoethanol or guanidine hydrochloride. Expression of the Hmd gene in E. coli without the co-factor results in an inactive holoenzyme. However hydrogenase activity can be rescued by the addition of the iron-containing co-factor taken from denatured active enzyme.[1]
As mentioned, irradiation of the co-factor with UV light results in the loss of CO and Fe. In addition the 542 Da compound can be further degraded by a phosphodiesterase (which specifically cleaves phosphate bonds). Hydrolysis of the phosphate bonds generates the ribonucleotide guanine mono-phosphate and a modified 2-pyridone. On the basis of spectroscopic characterization, Shima et al. have proposed a structure for this organic cofactor (minus the iron atom and CO molecules) as shown:[4]
Although the mechanism by which Hmd acts is unknown, the iron-containing cofactor is in part responsible for the catalytic activity. High concentrations of CO inhibit the enzyme as well, implicating iron as the center of catalysis. It has been proposed that the iron functions to bind H2 and the substrate methenyltetrahydromethanopterin, organizing these two reactants in close proximity.[1]
References
- Shima S, Thauer RK (2007). "A third type of hydrogenase catalyzing H2 activation". Chem Rec. 7 (1): 37–46. doi:10.1002/tcr.20111. PMID 17304591.
- "EC 1.12.98.2 - 5,10-methenyltetrahydromethanopterin hydrogenase". BRENDA – The comprehensive Enzyme Information System. Technische Universität Braunschweig.
- Lyon EJ, Shima S, Buurman G, Chowdhuri S, Batschauer A, Steinbach K, Thauer RK (January 2004). "UV-A/blue-light inactivation of the 'metal-free' hydrogenase (Hmd) from methanogenic archaea". Eur. J. Biochem. 271 (1): 195–204. doi:10.1046/j.1432-1033.2003.03920.x. PMID 14686932.
- Shima S, Lyon EJ, Sordel-Klippert M, Kauss M, Kahnt J, Thauer RK, Steinbach K, Xie X, Verdier L, Griesinger C (May 2004). "The cofactor of the iron-sulfur cluster free hydrogenase hmd: structure of the light-inactivation product". Angew. Chem. Int. Ed. Engl. 43 (19): 2547–51. doi:10.1002/anie.200353763. PMID 15127449.
Further reading
- Blaut M (1994). "Metabolism of methanogens". Antonie van Leeuwenhoek. 66 (1–3): 187–208. doi:10.1007/BF00871639. PMID 7747931.
- Karyakin AA, Varfolomeev SD (1986). "Catalytic Properties of Hydrogenases". Russian Chemical Reviews. 55: 867–882. doi:10.1070/rc1986v055n09abeh003228.