Pilin
Pilin refers to a class of fibrous proteins that are found in pilus structures in bacteria. Bacterial pili are used in the exchange of genetic material during bacterial conjugation, while a shorter type of appendages also made up of pilin, called fimbriae, are used as a cell adhesion mechanism. Although not all bacteria have pili or fimbriae, bacterial pathogens often use their fimbriae to attach to host cells. In Gram-negative bacteria, where pili are more common, individual pilin molecules are linked by noncovalent protein-protein interactions, while Gram-positive bacteria often have polymerized pilin.[1]
| Pilin (bacterial filament) | |||||||||
|---|---|---|---|---|---|---|---|---|---|
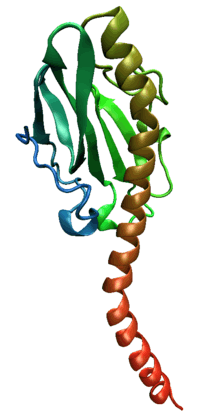 Pilin protein from Neisseria gonorrhoeae, a parasitic bacterium that requires functional pili for pathogenesis. | |||||||||
| Identifiers | |||||||||
| Symbol | Pili | ||||||||
| Pfam | PF00114 | ||||||||
| InterPro | IPR001082 | ||||||||
| PROSITE | PDOC00342 | ||||||||
| SCOPe | 1paj / SUPFAM | ||||||||
| OPM superfamily | 68 | ||||||||
| OPM protein | 2hil | ||||||||
| |||||||||
Some pilin proteins are α+β proteins characterized by a very long N-terminal alpha helix. The assembly of these pili relies on interactions between the N-terminal helices of the individual monomers. The pilus structure sequesters the helices in the center of the fiber lining a central pore, while antiparallel beta sheets occupy the exterior of the fiber.[2] The exact mechanism of assembly of these pili from monomers is not known, although chaperone proteins have been identified for some types of pilin.[3] and specific amino acids required for proper pilus formation have been isolated.[4]
Development of molecular tools
Pili in Gram-positive bacteria contain spontaneously formed isopeptide bonds. These bonds provide enhanced mechanical[5] and proteolytic[6] stability to the pilin protein. Recently, the pilin protein from Streptococcus pyogenes has been split into two fragments to develop a new molecular tool called the isopeptag.[7] The isopeptag is a short peptide that can be attached to a protein of interest and can bind its binding partner through a spontaneously formed isopeptide bond. This new peptide tag can allow scientists to target and isolate their proteins of interest through a permanent covalent bond.
Role of ComP pilin in bacterial transformation
Genetic transformation is the process by which a recipient bacterial cell takes up DNA from a neighboring cell and integrates this DNA into the recipient’s genome by homologous recombination. In Neisseria meningitidis, DNA transformation requires the presence of short DNA uptake sequences (DUSs) which are 9-10mers residing in coding regions of the donor DNA. Specific recognition of DUSs is mediated by a type IV pilin, ComP.[8][9] Menningococcal type IV pili bind DNA through the minor pilin ComP via an electropositive stripe that is predicted to be exposed on the filament's surface. ComP displays an exquisite binding preference for selective DUSs. The distribution of DUSs within the N. meningitidis genome favors certain genes, suggesting that there is a bias for genes involved in genomic maintenance and repair.[10][11]
See also
References
- Telford JL, Barocchi MA, Margarit I, Rappuoli R, Grandi G (2006). "Pili in gram-positive pathogens". Nat. Rev. Microbiol. 4 (7): 509–19. doi:10.1038/nrmicro1443. PMID 16778837.
- Forest KT, Tainer JA (1997). "Type-4 pilus-structure: outside to inside and top to bottom--a minireview". Gene. 192 (1): 165–9. doi:10.1016/s0378-1119(97)00008-5. PMID 9224887.
- Jones CH, Pinkner JS, Nicholes AV, Slonim LN, Abraham SN, Hultgren SJ (1993). "FimC is a periplasmic PapD-like chaperone that directs assembly of type 1 pili in bacteria". Proc. Natl. Acad. Sci. U.S.A. 90 (18): 8397–401. Bibcode:1993PNAS...90.8397J. doi:10.1073/pnas.90.18.8397. PMC 47363. PMID 8104335.
- Mu XQ, Jiang ZG, Bullitt E (2005). "Localization of a critical interface for helical rod formation of bacterial adhesion P-pili". J. Mol. Biol. 346 (1): 13–20. doi:10.1016/j.jmb.2004.11.037. PMID 15663923.
- Alegre-Cebollada J, Badilla CL, Fernández JM (2010). "Isopeptide bonds block the mechanical extension of pili in pathogenic Streptococcus pyogenes". J. Biol. Chem. 285 (15): 11235–11242. doi:10.1074/jbc.M110.102962. PMC 2857001. PMID 20139067.
- Kang HJ, Coulibaly F, Clow F, Proft T, Baker EN (2007). "Stabilizing isopeptide bonds revealed in gram-positive bacterial pilus structure". Science. 318 (5856): 1625–1628. Bibcode:2007Sci...318.1625K. doi:10.1126/science.1145806. PMID 18063798.
- Zakeri B, Howarth M (2010). "Spontaneous intermolecular amide bond formation between side chains for irreversible peptide targeting". J. Am. Chem. Soc. 132 (13): 4526–7. CiteSeerX 10.1.1.706.4839. doi:10.1021/ja910795a. PMID 20235501.
- Berry JL, Cehovin A, McDowell MA, Lea SM, Pelicic V (2013). "Functional analysis of the interdependence between DNA uptake sequence and its cognate ComP receptor during natural transformation in Neisseria species". PLOS Genet. 9 (12): e1004014. doi:10.1371/journal.pgen.1004014. PMC 3868556. PMID 24385921.
- Cehovin A, Simpson PJ, McDowell MA, Brown DR, Noschese R, Pallett M, Brady J, Baldwin GS, Lea SM, Matthews SJ, Pelicic V (2013). "Specific DNA recognition mediated by a type IV pilin". Proc. Natl. Acad. Sci. U.S.A. 110 (8): 3065–70. Bibcode:2013PNAS..110.3065C. doi:10.1073/pnas.1218832110. PMC 3581936. PMID 23386723.
- Davidsen T, Rødland EA, Lagesen K, Seeberg E, Rognes T, Tønjum T (2004). "Biased distribution of DNA uptake sequences towards genome maintenance genes". Nucleic Acids Res. 32 (3): 1050–8. doi:10.1093/nar/gkh255. PMC 373393. PMID 14960717.
- Caugant DA, Maiden MC (2009). "Meningococcal carriage and disease--population biology and evolution". Vaccine. 27 Suppl 2: B64–70. doi:10.1016/j.vaccine.2009.04.061. PMC 2719693. PMID 19464092.