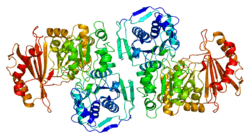PGM1
Phosphoglucomutase-1 is an enzyme that in humans is encoded by the PGM1 gene.[5][6][7] The protein encoded by this gene is an isozyme of phosphoglucomutase (PGM) and belongs to the phosphohexose mutase family. There are several PGM isozymes, which are encoded by different genes and catalyze the transfer of phosphate between the 1 and 6 positions of glucose. In most cell types, this PGM isozyme is predominant, representing about 90% of total PGM activity. In red blood cells, PGM2 is a major isozyme. This gene is highly polymorphic. Mutations in this gene cause CDG syndrome type 1t (CDG1T, formerly known as glycogen storage disease type XIV). Alternatively spliced transcript variants encoding different isoforms have been identified in this gene.[provided by RefSeq, Mar 2010][7]
Structure
The PGM1 gene is localized to the first chromosome, with its specific region being 1p31.[8] The complete PGM1 gene spans over 65 kb and contains 11 exons, and the sites of the two mutations which form the molecular basis for the common PGM1 protein polymorphism lie in exons 4 and 8 and are 18 kb apart. Within this region there is a site of intragenic recombination. There are two alternatively spliced first exons, one of which, exon 1A, is transcribed in a wide variety of cell types; the other, exon 1B, is transcribed in fast muscle tissue. Exon 1A is transcribed from a promoter that has the structural hallmarks of a housekeeping promoter but lies more than 35 kb upstream of exon 2. Exon 1B lies 6 kb upstream of exon 2 within the large first intron of the ubiquitously expressed PGM1 transcript. The fast-muscle form of PGM1 is characterized by 18 extra amino acid residues at its N-terminal end. Sequence comparisons show that exons 1A and 1B are structurally related and have arisen by duplication. [9]
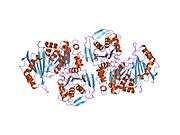
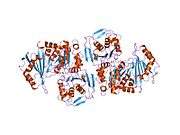
PGM1 is a monomeric protein with 562 amino acids and four structural domains arranged in an overall heart shape. The active site is located in the large, centrally located cleft, formed by more than 80 residues. The active site can be segregated into four highly conserved regions that contribute to catalysis and substrate binding.[10] These regions are: the phosphoserine residue that participates in phosphoryl transfer; the metal- binding loop; a sugar-binding loop; and the phosphate-binding site that interacts with the phosphate group of the substrate.[11] The active site cleft of PGM1 relies on all four structural domains of the enzyme for its structural integrity.[12][13]
Function
The biochemical pathways required to utilize glucose as a carbon and energy source are highly conserved from bacteria to humans. PGM1 is an evolutionarily conserved enzyme that regulates one of the most important metabolic carbohydrate trafficking points in prokaryotic and eukaryotic organisms, catalyzing the bi-directional interconversion of glucose 1-phosphate (G-1-P) and glucose 6-phosphate (G-6-P). In one direction, G-1-P produced from sucrose catabolism is converted to G-6-P, the first intermediate in glycolysis. In the other direction, conversion of G-6-P to G-1-P generates a substrate for synthesis of UDP-glucose, which is required for synthesis of a variety of cellular constituents, including cell wall polymers and glycoproteins.[14] PGM1 has been used extensively as a genetic marker for isozyme polymorphism among humans. PGM is known to be post-translationally modified by cytoplasmic glycosylation that does not seem to regulate its enzymatic activity but rather is implicated in the localization of the protein.[15] Glucose 1,6 bisphosphate (Glc-1, 6-P2), a powerful regulator of carbohydrate metabolism, has been demonstrated to be a potent activator of PGM. PGM1 is also modified by phosphorylation on Ser108 as part of its catalytic mechanism. This is shown to be performed by Pak1, a previously identified signaling kinase.[16]
Clinical significance
Phosphoglucomutase 1 (PGM1) deficiency is an inherited metabolic disorder in humans (CDG syndrome type 1t, CDG1T). Affected patients show multiple disease phenotypes, including dilated cardiomyopathy, exercise intolerance, and hepatopathy, reflecting the central role of the enzyme in glucose metabolism. The biochemical phenotypes of the PGM1 mutants cluster into two groups: those with compromised catalysis and those with possible folding defects. Relative to the recombinant wild-type enzyme, certain missense mutants show greatly decreased expression of soluble protein and/or increased aggregation. In contrast, other missense variants are well behaved in solution, but show dramatic reductions in enzyme activity, with Kcat/Km often <1.5% of wild-type. Modest changes in protein conformation and flexibility are also apparent in some of the catalytically impaired variants. In the case of the G291R mutant, severely compromised activity is linked to the inability of a key active site serine to be phosphorylated, a prerequisite for catalysis. Our results complement previous in vivo studies, which suggest that both protein misfolding and catalytic impairment may play a role in PGM1 deficiency.[17]
Interactions
PGM1 has been shown to interact with S100 calcium binding protein A1[18] and S100B.[18]
References
- GRCh38: Ensembl release 89: ENSG00000079739 - Ensembl, May 2017
- GRCm38: Ensembl release 89: ENSMUSG00000025791 - Ensembl, May 2017
- "Human PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- "Mouse PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- Douglas GR, McAlpine PJ, Hamerton JL (Oct 1973). "Regional localization of loci for human PGM and 6PGD on human chromosome one by use of hybrids of Chinese hamster-human somatic cells". Proceedings of the National Academy of Sciences of the United States of America. 70 (10): 2737–40. doi:10.1073/pnas.70.10.2737. PMC 427098. PMID 4517931.
- Whitehouse DB, Putt W, Lovegrove JU, Morrison K, Hollyoake M, Fox MF, Hopkinson DA, Edwards YH (Jan 1992). "Phosphoglucomutase 1: complete human and rabbit mRNA sequences and direct mapping of this highly polymorphic marker on human chromosome 1". Proceedings of the National Academy of Sciences of the United States of America. 89 (1): 411–5. doi:10.1073/pnas.89.1.411. PMC 48247. PMID 1530890.
- "Entrez Gene: PGM1 phosphoglucomutase 1".
- Douglas GR, McAlpine PJ, Hamerton JL (Oct 1973). "Regional localization of loci for human PGM and 6PGD on human chromosome one by use of hybrids of Chinese hamster-human somatic cells". Proceedings of the National Academy of Sciences of the United States of America. 70 (10): 2737–40. doi:10.1073/pnas.70.10.2737. PMC 427098. PMID 4517931.
- Putt W, Ives JH, Hollyoake M, Hopkinson DA, Whitehouse DB, Edwards YH (Dec 1993). "Phosphoglucomutase 1: a gene with two promoters and a duplicated first exon". The Biochemical Journal. 296 (2): 417–22. doi:10.1042/bj2960417. PMC 1137712. PMID 8257433.
- Shackelford GS, Regni CA, Beamer LJ (Aug 2004). "Evolutionary trace analysis of the alpha-D-phosphohexomutase superfamily". Protein Science. 13 (8): 2130–8. doi:10.1110/ps.04801104. PMC 2279825. PMID 15238632.
- Liu Y, Ray WJ, Baranidharan S (Jul 1997). "Structure of rabbit muscle phosphoglucomutase refined at 2.4 A resolution". Acta Crystallographica Section D. 53 (Pt 4): 392–405. doi:10.1107/S0907444997000875. PMID 15299905.
- Beamer LJ (Mar 2015). "Mutations in hereditary phosphoglucomutase 1 deficiency map to key regions of enzyme structure and function". Journal of Inherited Metabolic Disease. 38 (2): 243–56. doi:10.1007/s10545-014-9757-9. PMID 25168163.
- Luebbering EK, Mick J, Singh RK, Tanner JJ, Mehra-Chaudhary R, Beamer LJ (Nov 2012). "Conservation of functionally important global motions in an enzyme superfamily across varying quaternary structures". Journal of Molecular Biology. 423 (5): 831–46. doi:10.1016/j.jmb.2012.08.013. PMID 22935436.
- Boros LG, Lee WN, Go VL (Jan 2002). "A metabolic hypothesis of cell growth and death in pancreatic cancer". Pancreas. 24 (1): 26–33. CiteSeerX 10.1.1.537.3798. doi:10.1097/00006676-200201000-00004. PMID 11741179.
- Dey NB, Bounelis P, Fritz TA, Bedwell DM, Marchase RB (Oct 1994). "The glycosylation of phosphoglucomutase is modulated by carbon source and heat shock in Saccharomyces cerevisiae". The Journal of Biological Chemistry. 269 (43): 27143–8. PMID 7929458.
- Gururaj A, Barnes CJ, Vadlamudi RK, Kumar R (Oct 2004). "Regulation of phosphoglucomutase 1 phosphorylation and activity by a signaling kinase". Oncogene. 23 (49): 8118–27. doi:10.1038/sj.onc.1207969. PMID 15378030.
- Lee Y, Stiers KM, Kain BN, Beamer LJ (Nov 2014). "Compromised catalysis and potential folding defects in in vitro studies of missense mutants associated with hereditary phosphoglucomutase 1 deficiency". The Journal of Biological Chemistry. 289 (46): 32010–9. doi:10.1074/jbc.M114.597914. PMC 4231678. PMID 25288802.
- Landar A, Caddell G, Chessher J, Zimmer DB (Sep 1996). "Identification of an S100A1/S100B target protein: phosphoglucomutase". Cell Calcium. 20 (3): 279–85. doi:10.1016/S0143-4160(96)90033-0. PMID 8894274.
Further reading
- Dawson SJ, White LA (May 1992). "Treatment of Haemophilus aphrophilus endocarditis with ciprofloxacin". The Journal of Infection. 24 (3): 317–20. doi:10.1016/S0163-4453(05)80037-4. PMID 1602151.
- Herbich J, Szilvassy J, Schnedl W (1985). "Gene localisation of the PGM1 enzyme system and the Duffy blood groups on chromosome No. 1 by means of a new fragile site at 1p31". Human Genetics. 70 (2): 178–80. doi:10.1007/BF00273078. PMID 3159642.
- Takahashi N, Neel JV (Nov 1993). "Intragenic recombination at the human phosphoglucomutase 1 locus: predictions fulfilled". Proceedings of the National Academy of Sciences of the United States of America. 90 (22): 10725–9. doi:10.1073/pnas.90.22.10725. PMC 47850. PMID 7902567.
- March RE, Putt W, Hollyoake M, Ives JH, Lovegrove JU, Hopkinson DA, Edwards YH, Whitehouse DB (Nov 1993). "The classical human phosphoglucomutase (PGM1) isozyme polymorphism is generated by intragenic recombination". Proceedings of the National Academy of Sciences of the United States of America. 90 (22): 10730–3. doi:10.1073/pnas.90.22.10730. PMC 47851. PMID 7902568.
- Putt W, Ives JH, Hollyoake M, Hopkinson DA, Whitehouse DB, Edwards YH (Dec 1993). "Phosphoglucomutase 1: a gene with two promoters and a duplicated first exon". The Biochemical Journal. 296. 296 (2): 417–22. doi:10.1042/bj2960417. PMC 1137712. PMID 8257433.
- Edwards YH, Putt W, Fox M, Ives JH (Nov 1995). "A novel human phosphoglucomutase (PGM5) maps to the centromeric region of chromosome 9". Genomics. 30 (2): 350–3. doi:10.1006/geno.1995.9866. PMID 8586438.
- Moiseeva EP, Belkin AM, Spurr NK, Koteliansky VE, Critchley DR (Jan 1996). "A novel dystrophin/utrophin-associated protein is an enzymatically inactive member of the phosphoglucomutase superfamily". European Journal of Biochemistry / FEBS. 235 (1–2): 103–13. doi:10.1111/j.1432-1033.1996.00103.x. PMID 8631316.
- Landar A, Caddell G, Chessher J, Zimmer DB (Sep 1996). "Identification of an S100A1/S100B target protein: phosphoglucomutase". Cell Calcium. 20 (3): 279–85. doi:10.1016/S0143-4160(96)90033-0. PMID 8894274.
- Yip SP, Lovegrove JU, Rana NA, Hopkinson DA, Whitehouse DB (Sep 1999). "Mapping recombination hotspots in human phosphoglucomutase (PGM1)". Human Molecular Genetics. 8 (9): 1699–706. doi:10.1093/hmg/8.9.1699. PMID 10441333.
- Sergeev AS, Agapova RK, Bogadel'nikova IV, Perel'man MI (Jul 2003). "[The use of discrete characters in discriminant analysis for diagnosis of pulmonary tuberculosis and for classification of patients differing in treatment efficiency based on polymorphisms at nine codominant loci-HP, GC, TF, PI, PGM1, GLO1, C3, ACP1 and ESD]". Genetika. 39 (7): 996–1002. PMID 12942785.
- Gururaj A, Barnes CJ, Vadlamudi RK, Kumar R (Oct 2004). "Regulation of phosphoglucomutase 1 phosphorylation and activity by a signaling kinase". Oncogene. 23 (49): 8118–27. doi:10.1038/sj.onc.1207969. PMID 15378030.
- Takahashi K, Inuzuka M, Ingi T (Dec 2004). "Cellular signaling mediated by calphoglin-induced activation of IPP and PGM". Biochemical and Biophysical Research Communications. 325 (1): 203–14. doi:10.1016/j.bbrc.2004.10.021. PMID 15522220.
- Rual JF, Venkatesan K, Hao T, Hirozane-Kishikawa T, Dricot A, Li N, Berriz GF, Gibbons FD, Dreze M, Ayivi-Guedehoussou N, Klitgord N, Simon C, Boxem M, Milstein S, Rosenberg J, Goldberg DS, Zhang LV, Wong SL, Franklin G, Li S, Albala JS, Lim J, Fraughton C, Llamosas E, Cevik S, Bex C, Lamesch P, Sikorski RS, Vandenhaute J, Zoghbi HY, Smolyar A, Bosak S, Sequerra R, Doucette-Stamm L, Cusick ME, Hill DE, Roth FP, Vidal M (Oct 2005). "Towards a proteome-scale map of the human protein-protein interaction network". Nature. 437 (7062): 1173–8. doi:10.1038/nature04209. PMID 16189514.