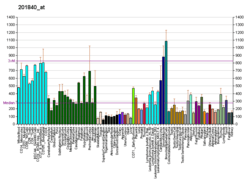
NEDD8
NEDD8 is a protein that in humans is encoded by the NEDD8 gene.[4][5] (In Saccharomyces cerevisiae this protein is known as Rub1.) This ubiquitin-like protein (ULP), becomes covalently conjugated to a limited number of cellular proteins in a manner analogous to ubiquitination. Human NEDD8 shares 60% amino acid sequence identity to ubiquitin. The primary known substrates of NEDD8 modification are the Cullin subunits of Cullin-based E3 ubiquitin ligases, which are active only when neddylated. Their NEDDylation is critical for the recruitment of E2 to the ligase complex, thus facilitating ubiquitin conjugation. NEDD8 modification has therefore been implicated in cell cycle progression and cytoskeletal regulation.
Activation and conjugation
As with ubiquitin and SUMO, NEDD8 is conjugated to cellular proteins after its C-terminal tail is processed. The NEDD8 activating E1 enzyme is a heterodimer composed of APPBP1 and UBA3 subunits.[6] The APPBP1/UBA3 enzyme has homology to the N- and C-terminal halves of the ubiquitin E1 enzyme, respectively. The UBA3 subunit contains the catalytic center and activates NEDD8 in an ATP-dependent reaction by forming a high-energy thiolester intermediate. The activated NEDD8 is subsequently transferred to the UbcH12 E2 enzyme, and is then conjugated to specific substrates in the presence of the appropriate E3 ligases.
Substrates for NEDD8
As reviewed by Brown et al.,[7] the best-characterized activated-NEDD8 substrates are the cullins (CUL1, 2, 3, 4A, 4B, 5, and 7 and PARC in human cells), that serve as molecular scaffolds for cullin-RING ubiquitin ligases (CRLs). Neddylation results in covalent conjugation of a NEDD8 moiety onto a conserved cullin lysine residue.[8] Cullin neddylation increases CRL ubiquitylation activity via conformational changes that optimize ubiquitin transfer to target proteins
Removal
There are several different proteases which can remove NEDD8 from protein conjugates. UCHL1, UCHL3 and USP21 proteases have dual specificity for NEDD8 and ubiquitin. Proteases specific for NEDD8 removal are the COP9 signalosome which removes NEDD8 from the CUL1 subunit of SCF ubiquitin ligases, and NEDP1 (or DEN1, SENP8).[9]
Role in DNA repair
As shown by Brown et al.,[7] NEDD8 accumulation at DNA-damage sites is a highly dynamic process. Neddylation is needed during a short period of the global genome repair (GGR) sub-pathway of DNA nucleotide excision repair (NER). In GGR of NER, after DNA damage is caused by UV irradiation, Cul4A in the DNA damage binding protein 2 (DDB2) complex is activated by NEDD8, and this allows GGR-NER to proceed to remove the damage.[10]
Neddylation also has a role in repair of double-strand breaks.[7] Non-homologous end joining(NHEJ) is a DNA repair pathway frequently used to repair DNA double-strand breaks. The first step in this pathway depends on the Ku70/Ku80 heterodimer that forms a highly stable ring structure encircling DNA ends.[11] But the Ku heterodimer needs to be removed when NHEJ is completed, or it blocks transcription or replication. The Ku heterodimer is ubiquitylated in a DNA-damage and neddylation-dependent manner to promote the release of Ku and other NHEJ factors from the site of repair after the process is completed.[7]
In cancer chemotherapy
As discussed by Jin and Roberston in their review,[12] silencing of a DNA repair gene by hypermethylation of its promoter may be a very early step in progression to cancer. Gene silencing of a DNA repair gene at the transcription level is proposed to act similarly to a germ-line mutation in a DNA repair gene. Loss of DNA repair capability by either mechanism introduces genome instability and predisposes the cell and its descendants to progression to cancer. Epigenetically silenced DNA repair genes occur frequently in the 17 most common cancers (see e.g. Frequency of hypermethylation of DNA repair genes in cancer).[12]
As discussed above, activated-NEDD8 is needed in two DNA repair pathways: NER and NHEJ. If activation of NEDD8 is inhibited, cells with induced deficiency of NER or NHEJ may then die because of deficient DNA repair leading to accumulation of DNA damages. The effect of NEDD8 inhibition may be greater for cancer cells than for normal cells if the cancer cells are independently deficient in DNA repair due to prior epigenetic silencing of DNA repair genes active in alternative pathways (see synthetic lethality).
Pevonedistat (MLN4924), a drug inhibiting activation of NEDD8, has shown a significant therapeutic effect in four Phase I clinical cancer trials in 2015-2016. These include pevonedistat trials against acute myeloid leukemia and myelodysplastic syndromes,[13] relapsed/refractory multiple myeloma or lymphoma,[14] metastatic melanoma,[15] and advanced solid tumors.[16]
In preclinical studies
PPARγ neddylation
PPARγ has a crucial role in adipogenesis and lipid accumulation within adipocytes (fat cells).[17] Activated NEDD8 stabilizes PPARγ, allowing increased adipogenesis. In experiments with mice, Pevonedistat, a drug inhibiting activation of NEDD8, prevented high-fat diet-induced obesity and glucose intolerance.[17]
NF-κB and NEDD8
The transcriptional activity of NF-κB is primarily regulated by physical interaction with inhibitory IκB proteins (IκBα and IκBβ), which prevents its nuclear translocation.[18] Degradation of the IκBα subunit of IκB is mediated by ubiquitination, and this ubiquitination depends on neddylation.[19] Pevonedistat (MLN4924) inhibits activation of NEDD8, that then inhibits ubiquitination of IκBα, and this inhibits NF-κB translocation to the nucleus.[18]
Pevonedistat, through its effects on NF-κB and a target of NF-κB (microRNA-155), prolonged the survival of mice engrafted with leukemic cells.[18]
Colorectal cancer
Inhibition of NEDD8 activation by pevonedistat was found to induce growth arrest and apoptosis in 16/122 (13%) colorectal cancer (CRC) cell lines. Further analyses in patient-derived tumor xenografts revealed that pevonedistat is effective on poorly differentiated, high-grade mucinous CRC. [20]
Interactions
NEDD8 has been shown to interact with:
References
- ENSG00000285246 GRCh38: Ensembl release 89: ENSG00000129559, ENSG00000285246 - Ensembl, May 2017
- "Human PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- "Mouse PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- Kamitani T, Kito K, Nguyen HP, Yeh ET (November 1997). "Characterization of NEDD8, a developmentally down-regulated ubiquitin-like protein". The Journal of Biological Chemistry. 272 (45): 28557–62. doi:10.1074/jbc.272.45.28557. PMID 9353319.
- "Entrez Gene: NEDD8 neural precursor cell expressed, developmentally down-regulated 8".
- Walden H, Podgorski MS, Huang DT, Miller DW, Howard RJ, Minor DL, Holton JM, Schulman BA (December 2003). "The structure of the APPBP1-UBA3-NEDD8-ATP complex reveals the basis for selective ubiquitin-like protein activation by an E1". Molecular Cell. 12 (6): 1427–37. doi:10.1016/s1097-2765(03)00452-0. PMID 14690597.
- Brown JS, Lukashchuk N, Sczaniecka-Clift M, Britton S, le Sage C, Calsou P, Beli P, Galanty Y, Jackson SP (May 2015). "Neddylation promotes ubiquitylation and release of Ku from DNA-damage sites". Cell Reports. 11 (5): 704–14. doi:10.1016/j.celrep.2015.03.058. PMC 4431666. PMID 25921528.
- Pan ZQ, Kentsis A, Dias DC, Yamoah K, Wu K (March 2004). "Nedd8 on cullin: building an expressway to protein destruction". Oncogene. 23 (11): 1985–97. doi:10.1038/sj.onc.1207414. PMID 15021886.
- "Boston Biochem NEDD8 Reagents Overview". Archived from the original on 2008-05-02. Retrieved 2008-04-29.
- Groisman R, Polanowska J, Kuraoka I, Sawada J, Saijo M, Drapkin R, Kisselev AF, Tanaka K, Nakatani Y (May 2003). "The ubiquitin ligase activity in the DDB2 and CSA complexes is differentially regulated by the COP9 signalosome in response to DNA damage". Cell. 113 (3): 357–67. doi:10.1016/s0092-8674(03)00316-7. PMID 12732143.
- Walker JR, Corpina RA, Goldberg J (August 2001). "Structure of the Ku heterodimer bound to DNA and its implications for double-strand break repair". Nature. 412 (6847): 607–14. Bibcode:2001Natur.412..607W. doi:10.1038/35088000. PMID 11493912.
- Jin B, Robertson KD (2013). DNA methyltransferases, DNA damage repair, and cancer. Advances in Experimental Medicine and Biology. 754. pp. 3–29. doi:10.1007/978-1-4419-9967-2_1. ISBN 978-1-4419-9966-5. PMC 3707278. PMID 22956494.
- Swords RT, Erba HP, DeAngelo DJ, Bixby DL, Altman JK, Maris M, Hua Z, Blakemore SJ, Faessel H, Sedarati F, Dezube BJ, Giles FJ, Medeiros BC (May 2015). "Pevonedistat (MLN4924), a First-in-Class NEDD8-activating enzyme inhibitor, in patients with acute myeloid leukaemia and myelodysplastic syndromes: a phase 1 study" (PDF). British Journal of Haematology. 169 (4): 534–43. doi:10.1111/bjh.13323. PMID 25733005.
- Shah JJ, Jakubowiak AJ, O'Connor OA, Orlowski RZ, Harvey RD, Smith MR, Lebovic D, Diefenbach C, Kelly K, Hua Z, Berger AJ, Mulligan G, Faessel HM, Tirrell S, Dezube BJ, Lonial S (January 2016). "Phase I Study of the Novel Investigational NEDD8-Activating Enzyme Inhibitor Pevonedistat (MLN4924) in Patients with Relapsed/Refractory Multiple Myeloma or Lymphoma". Clinical Cancer Research. 22 (1): 34–43. doi:10.1158/1078-0432.CCR-15-1237. PMC 5694347. PMID 26561559.
- Bhatia S, Pavlick AC, Boasberg P, Thompson JA, Mulligan G, Pickard MD, Faessel H, Dezube BJ, Hamid O (August 2016). "A phase I study of the investigational NEDD8-activating enzyme inhibitor pevonedistat (TAK-924/MLN4924) in patients with metastatic melanoma". Investigational New Drugs. 34 (4): 439–49. doi:10.1007/s10637-016-0348-5. PMC 4919369. PMID 27056178.
- Sarantopoulos J, Shapiro GI, Cohen RB, Clark JW, Kauh JS, Weiss GJ, Cleary JM, Mahalingam D, Pickard MD, Faessel HM, Berger AJ, Burke K, Mulligan G, Dezube BJ, Harvey RD (February 2016). "Phase I Study of the Investigational NEDD8-Activating Enzyme Inhibitor Pevonedistat (TAK-924/MLN4924) in Patients with Advanced Solid Tumors". Clinical Cancer Research. 22 (4): 847–57. doi:10.1158/1078-0432.CCR-15-1338. PMID 26423795.
- Park HS, Ju UI, Park JW, Song JY, Shin DH, Lee KH, Jeong LS, Yu J, Lee HW, Cho JY, Kim SY, Kim SW, Kim JB, Park KS, Chun YS (August 2016). "PPARγ neddylation essential for adipogenesis is a potential target for treating obesity". Cell Death and Differentiation. 23 (8): 1296–311. doi:10.1038/cdd.2016.6. PMC 4947677. PMID 26990658.
- Khalife J, Radomska HS, Santhanam R, Huang X, Neviani P, Saultz J, Wang H, Wu YZ, Alachkar H, Anghelina M, Dorrance A, Curfman J, Bloomfield CD, Medeiros BC, Perrotti D, Lee LJ, Lee RJ, Caligiuri MA, Pichiorri F, Croce CM, Garzon R, Guzman ML, Mendler JH, Marcucci G (October 2015). "Pharmacological targeting of miR-155 via the NEDD8-activating enzyme inhibitor MLN4924 (Pevonedistat) in FLT3-ITD acute myeloid leukemia". Leukemia. 29 (10): 1981–92. doi:10.1038/leu.2015.106. PMC 4868182. PMID 25971362.
- Frolova MA, Gudkova RG, Bol'shukhina LA, Novoselova VN (October 1978). "[Enhancement phenomenon in mice with a heterotopic heart transplant]". Zhurnal Mikrobiologii, Epidemiologii, I Immunobiologii. 20 (10): 36–40. PMID 85397.
- Picco G, Petti C, Sassi F, Grillone K, Migliardi G, Rossi T, Isella C, Di Nicolantonio F, Sarotto I, Sapino A, Bardelli A, Trusolino L, Bertotti A, Medico E (February 2017). "Efficacy of NEDD8 Pathway Inhibition in Preclinical Models of Poorly Differentiated, Clinically Aggressive Colorectal Cancer". Journal of the National Cancer Institute. 109 (2): djw209. doi:10.1093/jnci/djw209. PMID 27771609.
- Antenos M, Casper RF, Brown TJ (November 2002). "Interaction with Nedd8, a ubiquitin-like protein, enhances the transcriptional activity of the aryl hydrocarbon receptor". The Journal of Biological Chemistry. 277 (46): 44028–34. doi:10.1074/jbc.M202413200. PMID 12215427.
- Hipp MS, Raasi S, Groettrup M, Schmidtke G (April 2004). "NEDD8 ultimate buster-1L interacts with the ubiquitin-like protein FAT10 and accelerates its degradation". The Journal of Biological Chemistry. 279 (16): 16503–10. doi:10.1074/jbc.M310114200. PMID 14757770.
- Kamitani T, Kito K, Fukuda-Kamitani T, Yeh ET (December 2001). "Targeting of NEDD8 and its conjugates for proteasomal degradation by NUB1". The Journal of Biological Chemistry. 276 (49): 46655–60. doi:10.1074/jbc.M108636200. PMID 11585840.
- Gong L, Yeh ET (April 1999). "Identification of the activating and conjugating enzymes of the NEDD8 conjugation pathway". The Journal of Biological Chemistry. 274 (17): 12036–42. doi:10.1074/jbc.274.17.12036. PMID 10207026.
- Wada H, Kito K, Caskey LS, Yeh ET, Kamitani T (October 1998). "Cleavage of the C-terminus of NEDD8 by UCH-L3". Biochemical and Biophysical Research Communications. 251 (3): 688–92. doi:10.1006/bbrc.1998.9532. PMID 9790970.
Further reading
- Xirodimas DP (October 2008). "Novel substrates and functions for the ubiquitin-like molecule NEDD8". Biochemical Society Transactions. 36 (Pt 5): 802–6. doi:10.1042/BST0360802. PMID 18793140.
- Kumar S, Tomooka Y, Noda M (June 1992). "Identification of a set of genes with developmentally down-regulated expression in the mouse brain". Biochemical and Biophysical Research Communications. 185 (3): 1155–61. doi:10.1016/0006-291X(92)91747-E. PMID 1378265.
- Kumar S, Yoshida Y, Noda M (August 1993). "Cloning of a cDNA which encodes a novel ubiquitin-like protein". Biochemical and Biophysical Research Communications. 195 (1): 393–9. doi:10.1006/bbrc.1993.2056. PMID 8395831.
- Bonaldo MF, Lennon G, Soares MB (September 1996). "Normalization and subtraction: two approaches to facilitate gene discovery". Genome Research. 6 (9): 791–806. doi:10.1101/gr.6.9.791. PMID 8889548.
- Osaka F, Kawasaki H, Aida N, Saeki M, Chiba T, Kawashima S, Tanaka K, Kato S (August 1998). "A new NEDD8-ligating system for cullin-4A". Genes & Development. 12 (15): 2263–8. doi:10.1101/gad.12.15.2263. PMC 317039. PMID 9694792.
- Wada H, Kito K, Caskey LS, Yeh ET, Kamitani T (October 1998). "Cleavage of the C-terminus of NEDD8 by UCH-L3". Biochemical and Biophysical Research Communications. 251 (3): 688–92. doi:10.1006/bbrc.1998.9532. PMID 9790970.
- Whitby FG, Xia G, Pickart CM, Hill CP (December 1998). "Crystal structure of the human ubiquitin-like protein NEDD8 and interactions with ubiquitin pathway enzymes". The Journal of Biological Chemistry. 273 (52): 34983–91. doi:10.1074/jbc.273.52.34983. PMID 9857030.
- Gong L, Yeh ET (April 1999). "Identification of the activating and conjugating enzymes of the NEDD8 conjugation pathway". The Journal of Biological Chemistry. 274 (17): 12036–42. doi:10.1074/jbc.274.17.12036. PMID 10207026.
- Gong L, Li B, Millas S, Yeh ET (April 1999). "Molecular cloning and characterization of human AOS1 and UBA2, components of the sentrin-activating enzyme complex". FEBS Letters. 448 (1): 185–9. doi:10.1016/S0014-5793(99)00367-1. PMID 10217437.
- Liakopoulos D, Büsgen T, Brychzy A, Jentsch S, Pause A (May 1999). "Conjugation of the ubiquitin-like protein NEDD8 to cullin-2 is linked to von Hippel-Lindau tumor suppressor function". Proceedings of the National Academy of Sciences of the United States of America. 96 (10): 5510–5. Bibcode:1999PNAS...96.5510L. doi:10.1073/pnas.96.10.5510. PMC 21890. PMID 10318914.
- Singer JD, Gurian-West M, Clurman B, Roberts JM (September 1999). "Cullin-3 targets cyclin E for ubiquitination and controls S phase in mammalian cells". Genes & Development. 13 (18): 2375–87. doi:10.1101/gad.13.18.2375. PMC 317026. PMID 10500095.
- Simeoni S, Mancini MA, Stenoien DL, Marcelli M, Weigel NL, Zanisi M, Martini L, Poletti A (January 2000). "Motoneuronal cell death is not correlated with aggregate formation of androgen receptors containing an elongated polyglutamine tract" (PDF). Human Molecular Genetics. 9 (1): 133–44. doi:10.1093/hmg/9.1.133. PMID 10587588.
- Hori T, Osaka F, Chiba T, Miyamoto C, Okabayashi K, Shimbara N, Kato S, Tanaka K (November 1999). "Covalent modification of all members of human cullin family proteins by NEDD8". Oncogene. 18 (48): 6829–34. doi:10.1038/sj.onc.1203093. PMID 10597293.
- Read MA, Brownell JE, Gladysheva TB, Hottelet M, Parent LA, Coggins MB, Pierce JW, Podust VN, Luo RS, Chau V, Palombella VJ (April 2000). "Nedd8 modification of cul-1 activates SCF(beta(TrCP))-dependent ubiquitination of IkappaBalpha". Molecular and Cellular Biology. 20 (7): 2326–33. doi:10.1128/MCB.20.7.2326-2333.2000. PMC 85397. PMID 10713156.
- Morimoto M, Nishida T, Honda R, Yasuda H (April 2000). "Modification of cullin-1 by ubiquitin-like protein Nedd8 enhances the activity of SCF(skp2) toward p27(kip1)". Biochemical and Biophysical Research Communications. 270 (3): 1093–6. doi:10.1006/bbrc.2000.2576. PMID 10772955.
- Wada H, Yeh ET, Kamitani T (June 2000). "A dominant-negative UBC12 mutant sequesters NEDD8 and inhibits NEDD8 conjugation in vivo". The Journal of Biological Chemistry. 275 (22): 17008–15. doi:10.1074/jbc.275.22.17008. PMID 10828074.
- Kito K, Yeh ET, Kamitani T (June 2001). "NUB1, a NEDD8-interacting protein, is induced by interferon and down-regulates the NEDD8 expression". The Journal of Biological Chemistry. 276 (23): 20603–9. doi:10.1074/jbc.M100920200. PMID 11259415.
- Kamitani T, Kito K, Fukuda-Kamitani T, Yeh ET (December 2001). "Targeting of NEDD8 and its conjugates for proteasomal degradation by NUB1". The Journal of Biological Chemistry. 276 (49): 46655–60. doi:10.1074/jbc.M108636200. PMID 11585840.
- Wu K, Chen A, Tan P, Pan ZQ (January 2002). "The Nedd8-conjugated ROC1-CUL1 core ubiquitin ligase utilizes Nedd8 charged surface residues for efficient polyubiquitin chain assembly catalyzed by Cdc34". The Journal of Biological Chemistry. 277 (1): 516–27. doi:10.1074/jbc.M108008200. PMID 11675391.