HBx
HBx is a hepatitis B viral protein.[1][2] It is 154 amino acids long and interferes with transcription, signal transduction, cell cycle progress, protein degradation, apoptosis and chromosomal stability in the host. It forms a heterodimeric complex with its cellular target protein (HBX interacting protein: HBXIP), and this interaction dysregulates centrosome dynamics and mitotic spindle formation.[3] It interacts with DDB1 (Damaged DNA Binding Protein 1) redirecting the ubiquitin ligase activity of the CUL4-DDB1 E3 complexes, which are intimately involved in the intracellular regulation of DNA replication and repair, transcription and signal transduction.[4]
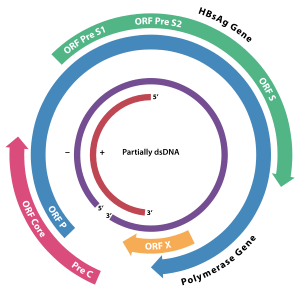
Although Protein X is normally absent in the Avihepadnavirus, a vestigial version has been identified in the duck hepatitis virus genome.[5]
Although it lacks significant sequence identity with any known vertebrate proteins, it seems likely that it evolved from a DNA glycosylase.[6]
Transgenic mice expressing the X protein in liver are more likely than the wild type to develop hepatocellular carcinoma. This is because the X protein promotes cell cycle progression while binding to and inhibiting tumor suppressor protein p53 from performing their role. Experimental observations also suggest that HBx protein increases TERT and telomerase activity, prolonging the lifespan of hepatocytes and contributing to malignant transformation.[7]
Molecular effects of HBx
HBx causes many cellular alterations. These alterations are due to direct actions of HBx and indirect actions due to large increases in intracellular reactive oxygen species (ROS) partly induced by HBx. HBx appears to dysregulate a number of cellular pathways. HBx causes dysregulation by binding to genomic DNA, changing expression patterns of miRNAs, affecting histone methyltransferases, binding to SIRT1 protein to activate transcription, and cooperating with histone methylases and demethylases to change cell expression patterns.[8]
HBx is partly responsible for the approximate 10,000-fold increase in intracellular ROS upon chronic HBV infection.[9][10] HBx can localize to the mitochondria where HBx decreases the mitochondrial membrane potential and causes increased release of ROS.[11] In addition, other HBV proteins, HBsAg[11] and HBcAg,[10] also increase ROS through interactions with the endoplasmic reticulum. ROS cause more than 20 types of DNA damage.[12] Oxidative DNA damage is mutagenic.[13]
HBx has large effects on the transcription levels of many genes. In a transgenic mouse model expressing the HBx gene of hepatitis B virus (but not other HBV genes), most mice developed hepatic tumors.[14] In these HBx transgenic mice there were 10,553 differentially DNA methylated regions (6,668 hypermethylated and 3,885 hypomethylated regions). In mammalian cells, large clusters of CpG dinucleotides known as CpG islands (CGIs) appear to act as key epigenetic elements regulating gene expression. Hyper-methylation of the CGIs in promoters can silence genes, while hypo-methylation of CGIs within distal exons of genes can also repress transcription of genes.[14] A large proportion of the methylation alterations in the HBx transgenic mice were at CGIs. HBx especially induced hypo-methylation of distal intragenic CGIs required for active expression. There were 647 genes containing intragenic CGIs that were hypo-methylated in HBx transgenic mouse liver.[14]
HBx also directly interacts with many genes. Several thousand protein-coding genes appear to have HBx-binding sites.[8][15] In addition to binding to protein coding genes, HBx bound to the promoters controlling 15 microRNAs and 16 Long non-coding RNAs.[15] For the 15 miRNAs with promoters bound by HBx, expression levels increased for eight, decreased for 5, and did not change significantly for two. Each microRNA with altered level of expression can affect the expression of several hundred messenger RNAs (see microRNA).
In addition to its effects on transcription levels of host genes, HBx seams to affect the in-vivo synthesis of pgRNA in HBV replicating cells. As HBx is recruited on cccDNA, it decreases levels of histone acetylation by decreasing recruitment p300 acetylases and increasing recruitment of hSirtl and HDAC1 deacetylases.[16] This, in turn, decreases heterochromatinization of HBV minichromosome, and increases the production of pgRNA. In cells infected by HBx defective mutants, levels of cccDNA remain unchanged, while there is a decrease in pgRNA transcription. Introduction of nucleus localized HBx protein seams to restore viral replication to cells infected by HBx-deficient virus.[17]
Relation to PRMT1
In a study purifying cancerous liver cells infected with HBV, the level of expression of protein arginine methyltransferase 1 (PRMT1) was found to be associated with changes in transcription due to the methyltransferase function of PRMT1. Overexpression causes a reduction in the number of HBV genes transcribed, while conversely, underexpression causes an increase. PRMT1 was also found to be recruited by HBV DNA during the replication process to regulate the transcription process. Increased HBx expression in turn leads to an inhibition of PRMT1-mediated protein methylation, benefiting viral replication.[18]
References
- McClain SL, Clippinger AJ, Lizzano R, Bouchard MJ (November 2007). "Hepatitis B virus replication is associated with an HBx-dependent mitochondrion-regulated increase in cytosolic calcium levels". Journal of Virology. 81 (21): 12061–5. doi:10.1128/JVI.00740-07. PMC 2168786. PMID 17699583.
- Bouchard MJ, Puro RJ, Wang L, Schneider RJ (July 2003). "Activation and inhibition of cellular calcium and tyrosine kinase signaling pathways identify targets of the HBx protein involved in hepatitis B virus replication". Journal of Virology. 77 (14): 7713–9. doi:10.1128/JVI.77.14.7713-7719.2003. PMC 161925. PMID 12829810.
- Wen Y, Golubkov VS, Strongin AY, Jiang W, Reed JC (February 2008). "Interaction of hepatitis B viral oncoprotein with cellular target HBXIP dysregulates centrosome dynamics and mitotic spindle formation". The Journal of Biological Chemistry. 283 (5): 2793–803. doi:10.1074/jbc.M708419200. PMID 18032378.
- Li T, Robert EI, van Breugel PC, Strubin M, Zheng N (January 2010). "A promiscuous alpha-helical motif anchors viral hijackers and substrate receptors to the CUL4-DDB1 ubiquitin ligase machinery". Nature Structural & Molecular Biology. 17 (1): 105–11. doi:10.1038/nsmb.1719. PMC 2823288. PMID 19966799.
- Lin B, Anderson DA (2000). "A vestigial X open reading frame in duck hepatitis B virus". Intervirology. 43 (3): 185–90. doi:10.1159/000025037. PMID 11044813.
- van Hemert FJ, van de Klundert MA, Lukashov VV, Kootstra NA, Berkhout B, Zaaijer HL (2011). "Protein X of hepatitis B virus: origin and structure similarity with the central domain of DNA glycosylase". PLOS ONE. 6 (8): e23392. Bibcode:2011PLoSO...623392V. doi:10.1371/journal.pone.0023392. PMC 3153941. PMID 21850270.
- Kew MC (January 2011). "Hepatitis B virus x protein in the pathogenesis of hepatitis B virus-induced hepatocellular carcinoma". Journal of Gastroenterology and Hepatology. 26 Suppl 1 (Suppl 1): 144–52. doi:10.1111/j.1440-1746.2010.06546.x. PMID 21199526.
- Balakrishnan L, Milavetz B (November 2017). "Epigenetic Regulation of Viral Biological Processes". Viruses. 9 (11): 346. doi:10.3390/v9110346. PMC 5707553. PMID 29149060.
- Valgimigli M, Valgimigli L, Trerè D, Gaiani S, Pedulli GF, Gramantieri L, Bolondi L (September 2002). "Oxidative stress EPR measurement in human liver by radical-probe technique. Correlation with etiology, histology and cell proliferation". Free Radical Research. 36 (9): 939–48. doi:10.1080/107156021000006653. PMID 12448819.
- Ivanov AV, Valuev-Elliston VT, Tyurina DA, Ivanova ON, Kochetkov SN, Bartosch B, Isaguliants MG (January 2017). "Oxidative stress, a trigger of hepatitis C and B virus-induced liver carcinogenesis". Oncotarget. 8 (3): 3895–3932. doi:10.18632/oncotarget.13904. PMC 5354803. PMID 27965466.
- Higgs MR, Chouteau P, Lerat H (May 2014). "'Liver let die': oxidative DNA damage and hepatotropic viruses" (PDF). The Journal of General Virology. 95 (Pt 5): 991–1004. doi:10.1099/vir.0.059485-0. PMID 24496828.
- Yu Y, Cui Y, Niedernhofer LJ, Wang Y (December 2016). "Occurrence, Biological Consequences, and Human Health Relevance of Oxidative Stress-Induced DNA Damage". Chemical Research in Toxicology. 29 (12): 2008–2039. doi:10.1021/acs.chemrestox.6b00265. PMC 5614522. PMID 27989142.
- Dizdaroglu M (December 2012). "Oxidatively induced DNA damage: mechanisms, repair and disease". Cancer Letters. 327 (1–2): 26–47. doi:10.1016/j.canlet.2012.01.016. PMID 22293091.
- Lee SM, Lee YG, Bae JB, Choi JK, Tayama C, Hata K, et al. (July 2014). "HBx induces hypomethylation of distal intragenic CpG islands required for active expression of developmental regulators". Proceedings of the National Academy of Sciences of the United States of America. 111 (26): 9555–60. Bibcode:2014PNAS..111.9555L. doi:10.1073/pnas.1400604111. PMC 4084425. PMID 24941955.
- Guerrieri F, Belloni L, D'Andrea D, Pediconi N, Le Pera L, Testoni B, et al. (February 2017). "Genome-wide identification of direct HBx genomic targets". BMC Genomics. 18 (1): 184. doi:10.1186/s12864-017-3561-5. PMC 5316204. PMID 28212627.
- Belloni L, Pollicino T, De Nicola F, Guerrieri F, Raffa G, Fanciulli M, et al. (November 2009). "Nuclear HBx binds the HBV minichromosome and modifies the epigenetic regulation of cccDNA function". Proceedings of the National Academy of Sciences of the United States of America. 106 (47): 19975–9. Bibcode:2009PNAS..10619975B. doi:10.1073/pnas.0908365106. PMC 2775998. PMID 19906987.
- Keasler VV, Hodgson AJ, Madden CR, Slagle BL (July 2009). "Hepatitis B virus HBx protein localized to the nucleus restores HBx-deficient virus replication in HepG2 cells and in vivo in hydrodynamically-injected mice". Virology. 390 (1): 122–9. doi:10.1016/j.virol.2009.05.001. PMC 2749295. PMID 19464721.
- Benhenda S, Ducroux A, Rivière L, Sobhian B, Ward MD, Dion S, et al. (April 2013). "Methyltransferase PRMT1 is a binding partner of HBx and a negative regulator of hepatitis B virus transcription". Journal of Virology. 87 (8): 4360–71. doi:10.1128/JVI.02574-12. PMC 3624337. PMID 23388725.