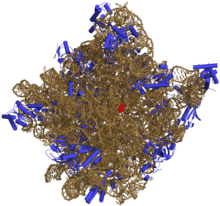
Prokaryotic large ribosomal subunit
50S is the larger subunit of the 70S ribosome of prokaryotes, i.e. bacteria and archaea. It is the site of inhibition for antibiotics such as macrolides, chloramphenicol, clindamycin, and the pleuromutilins. It includes the 5S ribosomal RNA and 23S ribosomal RNA.
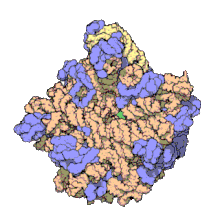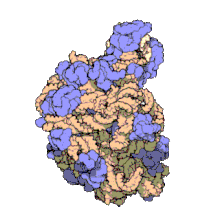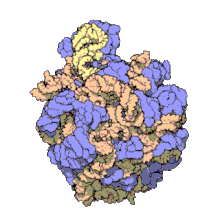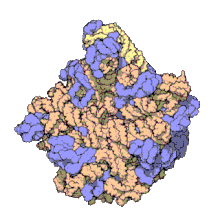
Structure
50S, roughly equivalent to the 60S ribosomal subunit in eukaryotic cells, is the larger subunit of the 70S ribosome of prokaryotes. The 50S subunit is primarily composed of proteins but also contains single-stranded RNA known as ribosomal RNA (rRNA). rRNA forms secondary and tertiary structures to maintain the structure and carry out the catalytic functions of the ribosome.
X-ray crystallography has yielded electron density maps allowing the structure of the 50S in Haloarcula marismortui to be determined to 2.4Å resolution[1]and of the 50S in Deinococcus radiodurans to 3.3Å.[2] The large ribosomal subunit (50S) is approximately twice as massive as the small ribosomal subunit (30S). The model of Hm 50S, determined in 2000 by Nenad Ban and colleagues in the laboratory of Thomas Steitz and the laboratory of Peter Moore, includes 2711 of the 2923 nucleotides of 23S rRNA, all 122 nucleotides of its 5S rRNA, and structure of 27 of its 31 proteins.[1]
A cryoEM structure of the 50S subunit from the archaeon Methanothermobacter thermautotrophicus has been determined. It shares the 50S size/sedimentation rate and the two rRNA count, but its 23S expansion segments have more in common with eukaryotes.[3]
Ribosomal RNA
The secondary structure of 23S is divided into six large domains, within which domain V is most important in its peptidyl transferase activity. Each domain contains normal secondary structure (e.g., base triple, tetraloop, cross-strand purine stack) and is also highly symmetric in tertiary structure; proteins intervene between their helices. At tertiary structure level, the large subunit rRNA is a single gigantic domain while the small subunit contains three structural domains. This difference reflects the lesser flexibility of the large subunit required by its function.
Function
50S includes the activity that catalyzes peptide bond formation (peptidyl transfer reaction), prevents premature polypeptide hydrolysis, provides a binding site for the G-protein factors (assists initiation, elongation, and termination), and helps protein folding after synthesis.
Promotes the peptidyl transfer reaction and prevents peptidyl hydrolysis
An induced-fit mechanism has been revealed for how 50S catalyzes the peptidyl transfer reaction and prevents peptidyl hydrolysis. The amino group of an aminoacyl-tRNA (binds to A site) attacks the carbon of a carbonyl group of a peptidyl-tRNA (binds to P site) and finally yields a peptide extended by one amino acid esterified to the A site tRNA bound to the ribosomal A site and a deacylated tRNA in the P site.
When the A site is unoccupied, nucleotide U2620 (E. coli U2585), A2486 (2451) and C2106 (2063) sandwich the carbonyl group in the middle, forcing it into an orientation facing the A site. This orientation prevents any nucleophilic attack from the A site because the optimal attacking angle is 105 degrees from the plane of the ester group. When a tRNA with a complete[?] CCA sequence at its acceptor stem is bound to the A site, C74 of the tRNA stacking with U2590 (2555) induces a conformational change in the ribosome, resulting in movement of U2541 (2506), U2620 (2585) through G2618 (2583). The displacement of bases allows the ester group to adopt a new conformation accessible to nucleophilic attack from the A site.
The N3 (nitrogen) of A2486 (2451) is closest to the peptide bond being synthesized and may function as a general base to facilitate the nucleophilic attack by the amino group of the aminoacyl-tRNA (in the A site). The pKa of A2486 (2451) is about 5 units higher in order to hydrogen bond with the amino group thus increasing its nucleophilicity. The elevation of pKa is achieved through a charge relay mechanism. A2486 (2451) interacts with G2482 (G2447), which hydrogen bonds with the buried phosphate of A2486 (2450). This buried phosphate can stabilize the normally rare imino tautomers of both bases, resulting in an increase in the negative charge density on N3.
Helps protein formation
After initiation, elongation, and termination, there is a fourth step of the disassembly of the post-termination complex of ribosome, mRNA, and tRNA, which is a prerequisite for the next round of protein synthesis. The large ribosomal subunit has a role in protein folding both in vitro and in vivo. The large ribosomal subunit provides a hydrophobic surface for the hydrophobic collapse step of protein folding. The newly synthesized protein needs full access to the large subunit to fold; this process may take a period of time (5 minutes for beta-galactosidase).
See also
- Prokaryotic small ribosomal subunit (30S)
- Ribosomal RNA
- 23S methyl RNA motif
References
- Nissen, P.; Hansen, J.; Ban, N.; Moore, P.; Steitz, T. (2000). "The complete atomic structure of the large ribosomal subunit at 2.4 A resolution". Science. 289 (5481): 905–920. CiteSeerX 10.1.1.58.2271. doi:10.1126/science.289.5481.905. PMID 10937989.
- Schluenzen, F.; Tocilj, A.; Zarivach, R.; Harms, J.; Gluehmann, M.; Janell, D.; Bashan, A.; Bartels, H.; Agmon, I.; Franceschi, F.; Yonath, A. (2000). "Structure of functionally activated small ribosomal subunit at 3.3 Å resolution". Cell. 102 (5): 615–623. doi:10.1016/S0092-8674(00)00084-2. PMID 11007480.
- Greber, Basil J.; Boehringer, Daniel; Godinic-Mikulcic, Vlatka; Crnkovic, Ana; Ibba, Michael; Weygand-Durasevic, Ivana; Ban, Nenad (May 2012). "Cryo-EM Structure of the Archaeal 50S Ribosomal Subunit in Complex with Initiation Factor 6 and Implications for Ribosome Evolution". Journal of Molecular Biology. 418 (3–4): 145–160. doi:10.1016/j.jmb.2012.01.018.
- Nissen, P.; Hansen, J.; Ban, N.; Moore, P.; Steitz, T. (2000). "The structural basis of ribosome activity in peptide bond synthesis". Science. 289 (5481): 920–929.
- Schmeing, T.; Huang, K.; Strobel, S.; Steitz, T. (2005). "An induced-fit mechanism to promote peptide bond formation and exclude hydrolysis of peptidyl-tRNA". Nature. 438: 520–524.
- Basu, A.; Ghosh, J.; Bhattacharya, A.; Pal, S.; Chowdhury, S.; DasGupta, C. (2003). "Splitting of ribosome into its subunits by unfolded polypeptide chains". Current Science. 84: 1123–1125.
External links
- http://pathmicro.med.sc.edu/mayer/antibiot.htm
- https://web.archive.org/web/20110227235620/http://www.molgen.mpg.de/~ag_ribo/ag_franceschi/franceschi-projects-50S-antibiotics.html
- https://web.archive.org/web/20080206051722/http://www.riboworld.com/antib/50santib-eng.shtml
- 23S+Ribosomal+RNA at the US National Library of Medicine Medical Subject Headings (MeSH)
- 5S+Ribosomal+RNA at the US National Library of Medicine Medical Subject Headings (MeSH)