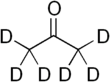
Deuterated acetone
Deuterated acetone ((CD3)2CO), also known as Acetone-D6, is a form (called an isotopologue) of acetone (CH3)2CO in which the hydrogen atom ("H") is replaced with deuterium (heavy hydrogen) isotope ("D"). Deuterated acetone is a common solvent used in NMR spectroscopy.
| |||
 | |||
| Identifiers | |||
|---|---|---|---|
3D model (JSmol) |
|||
| 1702935 | |||
| ChEBI | |||
| ChemSpider | |||
| ECHA InfoCard | 100.010.514 | ||
| EC Number |
| ||
PubChem CID |
|||
| UNII | |||
| UN number | 1090 | ||
CompTox Dashboard (EPA) |
|||
| |||
| |||
| Properties | |||
| C32H6O | |||
| Molar mass | 64.1161 g mol−1 | ||
| Density | 0.872 g cm−3 | ||
| Melting point | −94 °C (−137 °F; 179 K) | ||
| Boiling point | 56 °C (133 °F; 329 K) | ||
| Vapor pressure | 24.5-25.3 kPa (at 20 °C) | ||
| Hazards | |||
| GHS pictograms |   | ||
| GHS Signal word | Danger | ||
GHS hazard statements |
H225, H319, H336 | ||
| P210, P233, P240, P241, P242, P243, P261, P264, P271, P280, P303+361+353, P304+340, P305+351+338, P312, P337+313, P370+378, P403+233, P403+235, P405, P501 | |||
| NFPA 704 (fire diamond) | |||
| Flash point | −19 °C (−2 °F; 254 K) | ||
| Related compounds | |||
Related compounds |
Acetone | ||
Except where otherwise noted, data are given for materials in their standard state (at 25 °C [77 °F], 100 kPa). | |||
| Infobox references | |||
Properties
As with all deuterated compounds, the properties of deuterated acetone are virtually identical to those of regular acetone.
Manufacture
Deuterated acetone is prepared from heavy water, D2O, by what amounts to an aldol reaction. In this case, the base used is a deuterated version of lithium hydroxide:[1]
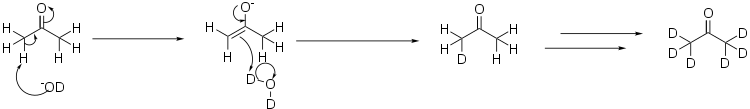

In order to fully deuterate the acetone, the process is repeated several times, distilling off the acetone from the heavy water, and re-running the reaction in a fresh batch of heavy water.
gollark: I also do not believe in the afterlife, but I am still against eternal torture abstractly speaking.
gollark: Also finite torture, in most cases.
gollark: I do not support eternal torture of any form.
gollark: Christianity's pretty bad too because it has hell, although *some* people argue you don't get eternal torture but just annihilated, which isn't much better, and also some people argue everyone goes to heaven or whatever because christianity is a mess.
gollark: Idea: omniquantism.
References
- P. J. Paulsen, W. D. Cooke. . Anal. Chem., 1963, 35 (10), pp 1560–1560. DOI: 10.1021/ac60203a072
This article is issued from Wikipedia. The text is licensed under Creative Commons - Attribution - Sharealike. Additional terms may apply for the media files.