XCMS Online
XCMS Online is bioinformatics software designed for statistical analysis of mass spectrometry data, created by the Siuzdak Lab at Scripps Research. It was designed to be an easy way to visualize and share untargeted metabolomic data.[1]
 | |
| Initial release | 25 April 2012 |
|---|---|
| Stable release | 3.7.1
|
| Platform | Open Web Platform |
| Type | Bioinformatics / Mass spectrometry software |
| License | Freemium / commercial software |
| Website | xcmsonline |
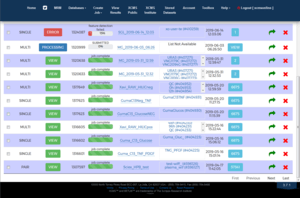
XCMS Online works by comparing groups of raw or preprocessed metabolomic data to discover metabolites using methods such as nonlinear retention time alignment and feature detection & matching. Once analysis is complete the data can be viewed several different ways including via bubble plots, heat maps, chromatograms, and box plots. In addition, XCMS Online is integrated with METLIN, a large metabolite database.[2][3] The following file formats are supported for direct upload to the site:[4]
| File Type | Vendor |
|---|---|
| mzXML | Open Format |
| mzData | Open Format |
| .cdf | NetCDF (AIA/ANDI) |
| .d folders | Agilent, Bruker |
| .wiff | SCIEX |
| .RAW folders | Waters |
| .RAW files | Thermo |
History
In 2005, the Siuzdak Lab created an open-source tool named XCMS in the programming language R. Noticing the need for a more accessible, graphical data processing tool they created the cloud-based XCMS Online in 2012.[1][5] The ability for users to stream data directly from instruments while being acquired was added in 2014.[6] Also in that year a commercial version named XCMS Plus (owned by Mass Consortium Corporation) was released and, in 2015, SCIEX became an exclusive reseller.[7] In 2017 it was shown that XCMS Online could be used in a systems biology workflow.[8] One year later, in the absence of a publicly available alternative, a version of XCMS Online was released with the ability to perform multiple reaction monitoring (MRM).[9]
References
- Tautenhahn, Ralf; Patti, Gary J.; Rinehart, Duane; Siuzdak, Gary (5 June 2012). "XCMS Online: A Web-Based Platform to Process Untargeted Metabolomic Data". Analytical Chemistry. 84 (11): 5035–5039. doi:10.1021/ac300698c. PMC 3703953.
- Smith, Colin A.; Want, Elizabeth J.; O'Maille, Grace; Abagyan, Ruben; Siuzdak, Gary (February 2006). "XCMS: Processing Mass Spectrometry Data for Metabolite Profiling Using Nonlinear Peak Alignment, Matching, and Identification". Analytical Chemistry. 78 (3): 779–787. doi:10.1021/ac051437y.
- Gowda, Harsha; Ivanisevic, Julijana; Johnson, Caroline H.; Kurczy, Michael E.; Benton, H. Paul; Rinehart, Duane; Nguyen, Thomas; Ray, Jayashree; Kuehl, Jennifer; Arevalo, Bernardo; Westenskow, Peter D.; Wang, Junhua; Arkin, Adam P.; Deutschbauer, Adam M.; Patti, Gary J.; Siuzdak, Gary (25 June 2014). "Interactive XCMS Online: Simplifying Advanced Metabolomic Data Processing and Subsequent Statistical Analyses". Analytical Chemistry. 86 (14): 6931–6939. doi:10.1021/ac500734c. PMC 4215863.
- "XCMS Online - Documentation". xcmsonline.scripps.edu.
- Perkel, Jeffrey M. "Name That Metabolite!". The Scientist Magazine®. The Scientist. Retrieved 25 June 2019.
- Rinehart, Duane; Johnson, Caroline H; Nguyen, Thomas; Ivanisevic, Julijana; Benton, H Paul; Lloyd, Jessica; Arkin, Adam P; Deutschbauer, Adam M; Patti, Gary J; Siuzdak, Gary (9 June 2014). "Metabolomic data streaming for biology-dependent data acquisition". Nature Biotechnology. 32 (6): 524–527. doi:10.1038/nbt.2927. PMC 4112958.
- Evans, Jon. "SCIEX becomes exclusive reseller for XCMS plus - News - spectroscopyNOW.com". www.spectroscopynow.com. Spectroscopy Now. Retrieved 25 June 2019.
- Huan, Tao; Forsberg, Erica M; Rinehart, Duane; Johnson, Caroline H; Ivanisevic, Julijana; Benton, H Paul; Fang, Mingliang; Aisporna, Aries; Hilmers, Brian; Poole, Farris L; Thorgersen, Michael P; Adams, Michael W W; Krantz, Gregory; Fields, Matthew W; Robbins, Paul D; Niedernhofer, Laura J; Ideker, Trey; Majumder, Erica L; Wall, Judy D; Rattray, Nicholas J W; Goodacre, Royston; Lairson, Luke L; Siuzdak, Gary (1 May 2017). "Systems biology guided by XCMS Online metabolomics". Nature Methods. 14 (5): 461–462. doi:10.1038/nmeth.4260. PMC 5933448.
- Domingo-Almenara, Xavier; Montenegro-Burke, J. Rafael; Ivanisevic, Julijana; Thomas, Aurelien; Sidibé, Jonathan; Teav, Tony; Guijas, Carlos; Aisporna, Aries E.; Rinehart, Duane; Hoang, Linh; Nordström, Anders; Gómez-Romero, María; Whiley, Luke; Lewis, Matthew R.; Nicholson, Jeremy K.; Benton, H. Paul; Siuzdak, Gary (27 August 2018). "XCMS-MRM and METLIN-MRM: a cloud library and public resource for targeted analysis of small molecules". Nature Methods. 15 (9): 681–684. doi:10.1038/s41592-018-0110-3. PMC 6629029.