Sirtuin 5
Sirtuin (silent mating type information regulation 2 homolog) 5 (S. cerevisiae), also known as SIRT5 is a protein which in humans in encoded by the SIRT5 gene and in other species by the orthologous Sirt5 gene.[5]
This gene encodes a member of the sirtuin family of proteins, homologs to the yeast Sir2 protein. Members of the sirtuin family are characterized by a sirtuin core domain and belong to the class III of the [histone deacetylase] superfamily, and are dependent on NAD+ as co-factor of enzymatic activities. SIRT5 is one of the three sirtuins localized primarily to the mitochondrion.
Structure
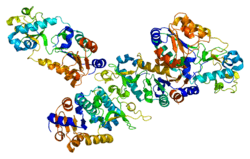
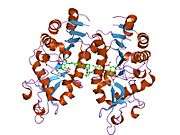
Alternative splicing of this gene results in two transcript variants.[5] The protein structure of SIRT5 has been resolved and shows high degrees of structural conservation with other sirtuins, such as the ancestral yeast protein and human SIRT2.
Function
SIRT5 has been found to exhibit enzymatic activities as a deacetylase, desuccinylase, and demalonylase, capable of removing acetyl, succinyl, and malonyl groups from the lysine residues of proteins.[6] SIRT5 deacetylases and regulates carbamoyl phosphate synthetase (CPS1), the rate-limiting and initiating step of the urea cycle in liver mitochondria. Deacetylation of CPS1 stimulates its enzymatic activity. Mice with deletion of SIRT5 show elevated ammonia levels after a prolonged fast, whereas in contrast, mice overexpressing SIRT5 show increased CPS1 activity, suggesting one of the functions of SIRT5 may be to regulate the urea cycle.[7] SIRT5 also interacts with and deacetylates cytochrome c.[8] Large-scale profiling studies of SIRT5 deacetylase activity have uncovered over 700 protein substrates, including proteins localized to the mitochondria, the cytosol and other sub cellular localization. The identities of SIRT5 desuccinylation substrates suggest that SIRT5-mediated desuccinylation may be involved in energy metabolism.[9]
The physiological consequences of SIRT5 molecular functions in human is under investigation but may involved regulations of mitochondrial metabolism.[10]
Interactions
Carbamoyl phosphate synthetase (CPS1)
References
- GRCh38: Ensembl release 89: ENSG00000124523 - Ensembl, May 2017
- GRCm38: Ensembl release 89: ENSMUSG00000054021 - Ensembl, May 2017
- "Human PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- "Mouse PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- "Entrez Gene: SIRT5 sirtuin (silent mating type information regulation 2 homolog) 5 (S. cerevisiae)".
- Du J, Zhou Y, Su X, Yu JJ, Khan S, Jiang H, Kim J, Woo J, Kim JH, Choi BH, He B, Chen W, Zhang S, Cerione RA, Auwerx J, Hao Q, Lin H (2011). "Sirt5 is a NAD-dependent protein lysine demalonylase and desuccinylase". Science. 334 (6057): 806–09. Bibcode:2011Sci...334..806D. doi:10.1126/science.1207861. PMC 3217313. PMID 22076378.
- Nakagawa T, Lomb DJ, Haigis MC, Guarente L (2009). "SIRT5 Deacetylates carbamoyl phosphate synthetase 1 and regulates the urea cycle". Cell. 137 (3): 560–70. doi:10.1016/j.cell.2009.02.026. PMC 2698666. PMID 19410549.
- Schlicker C, Gertz M, Papatheodorou P, Kachholz B, Becker CF, Steegborn C (2008). "Substrates and regulation mechanisms for the human mitochondrial sirtuins Sirt3 and Sirt5". J. Mol. Biol. 382 (3): 790–801. doi:10.1016/j.jmb.2008.07.048. PMID 18680753.
- Park J, Chen Y, Tishkoff DX, Peng C, Tan M, Dai L, Xie Z, Zhang Y, Zwaans BM, Skinner ME, Lombard DB, Zhao Y (2013). "SIRT5-mediated lysine desuccinylation impacts diverse metabolic pathways". Mol. Cell. 50 (6): 919–30. doi:10.1016/j.molcel.2013.06.001. PMC 3769971. PMID 23806337.
- Verdin E, Hirschey MD, Finley LW, Haigis MC (2010). "Sirtuin regulation of mitochondria: energy production, apoptosis, and signaling". Trends Biochem. Sci. 35 (12): 669–75. doi:10.1016/j.tibs.2010.07.003. PMC 2992946. PMID 20863707.
- Schlicker C, Gertz M, Papatheodorou P, Kachholz B, Becker CF, Steegborn C (2008). "Substrates and regulation mechanisms for the human mitochondrial sirtuins Sirt3 and Sirt5". J. Mol. Biol. 382 (3): 790–801. doi:10.1016/j.jmb.2008.07.048. PMID 18680753.
Further reading
- Frye RA (Jun 1999). "Characterization of five human cDNAs with homology to the yeast SIR2 gene: Sir2-like proteins (sirtuins) metabolize NAD and may have protein ADP-ribosyltransferase activity". Biochemical and Biophysical Research Communications. 260 (1): 273–79. doi:10.1006/bbrc.1999.0897. PMID 10381378.
- Frye RA (Jul 2000). "Phylogenetic classification of prokaryotic and eukaryotic Sir2-like proteins". Biochemical and Biophysical Research Communications. 273 (2): 793–98. doi:10.1006/bbrc.2000.3000. PMID 10873683.
- Hartley JL, Temple GF, Brasch MA (Nov 2000). "DNA cloning using in vitro site-specific recombination". Genome Research. 10 (11): 1788–95. doi:10.1101/gr.143000. PMC 310948. PMID 11076863.
- Wiemann S, Arlt D, Huber W, Wellenreuther R, Schleeger S, Mehrle A, Bechtel S, Sauermann M, Korf U, Pepperkok R, Sültmann H, Poustka A (Oct 2004). "From ORFeome to biology: a functional genomics pipeline". Genome Research. 14 (10B): 2136–44. doi:10.1101/gr.2576704. PMC 528930. PMID 15489336.
- Mehrle A, Rosenfelder H, Schupp I, del Val C, Arlt D, Hahne F, Bechtel S, Simpson J, Hofmann O, Hide W, Glatting KH, Huber W, Pepperkok R, Poustka A, Wiemann S (Jan 2006). "The LIFEdb database in 2006". Nucleic Acids Research. 34 (Database issue): D415–18. doi:10.1093/nar/gkj139. PMC 1347501. PMID 16381901.
- Mahlknecht U, Ho AD, Letzel S, Voelter-Mahlknecht S (2006). "Assignment of the NAD-dependent deacetylase sirtuin 5 gene (SIRT5) to human chromosome band 6p23 by in situ hybridization". Cytogenetic and Genome Research. 112 (3–4): 208–12. doi:10.1159/000089872. PMID 16484774.
- Chowdari KV, Northup A, Pless L, Wood J, Joo YH, Mirnics K, Lewis DA, Levitt PR, Bacanu SA, Nimgaonkar VL (Apr 2007). "DNA pooling: a comprehensive, multi-stage association analysis of ACSL6 and SIRT5 polymorphisms in schizophrenia". Genes, Brain, and Behavior. 6 (3): 229–39. doi:10.1111/j.1601-183X.2006.00251.x. PMID 16827919.