R2 RNA element
The R2 RNA element is a non-long terminal repeat (non-LTR) retrotransposable element that inserts at a specific site in the 28S ribosomal RNA (rRNA) genes of most insect genomes.[1] In order to insert itself into the genome, retrotransposon encoded protein (R2) protein makes a specific nick in one of the DNA strands at the insertion site and uses the 3′ hydroxyl group exposed by this nick to prime the reverse transcription process termed target primed reverse transcription (TPRT), where the RNA genome is transcribed into DNA.[2]
| R2 RNA element | |
|---|---|
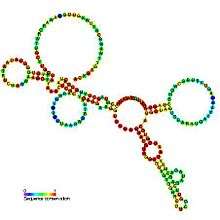 Predicted secondary structure and sequence conservation of R2_retro_el | |
| Identifiers | |
| Symbol | R2_retro_el |
| Rfam | RF00524 |
| Other data | |
| RNA type | Cis-reg |
| Domain(s) | Eukaryota |
| SO | 0000233 |
| PDB structures | PDBe |
3' UTR element
The R2 element 3' UTR RNA is a cis-acting element identified in R2 retrotransposons which is involved in priming the reverse transcription process (an essential part of retrotransposon insertion into the host genome).[3] An RNA fragment found in the R2 3' untranslated region (3'UTR), has been shown to interact with one copy of R2 protein during TPRT. This fragment has been shown to possess conserved secondary structure within Drosophila and silk moths, and also between the two groups.[3]
5' UTR ribozyme
The R2 element is co-transcribed with host organism 28S ribosomal RNA (rRNA). To become a fully mature R2 messenger RNA (mRNA), requires that the initial R2 transcript be processed to remove the 28S rRNA. This processing occurs via a self-cleaving ribozyme that forms at the 5' junction of the R2 RNA.[4] This ribozyme has been found to have high structural similarity to the HDV ribozyme but they are not homologous; the two sequences are thought to have undergone convergent evolution.[4]
The 5′ R2 protein binding site
The 5′ R2 protein binding site occurs in a region that spans part of the 5' UTR and the start of the R2 ORF. This region also has a conserved secondary structure, which has been deduced from binding to oligonucleotide microarrays, structure probing, and free energy minimization.[5] To date, conservation of structure has only been described in five moth species: Bombyx mori (R2Bm), Samia cynthia (R2Sc), Coscinocera hercules (R2Ch), Callosamia promethea (R2Cpr), Saturnia pyri (R2Spy)

Within this 5' protein binding site an RNA pseudoknot structure occurs.[6] The pseudoknot is highly conserved between the 5 silk moth species. Sequence comparisons show evidence for compensatory mutations within the helical regions indicating the secondary structure of the RNA is of biological importance. In particular, this pseudoknot is proposed to have implications for initiation of translation.[7]
References
- Luan DD, Korman MH, Jakubczak JL, Eickbush TH (1993). "Reverse transcription of R2Bm RNA is primed by a nick at the chromosomal target site: a mechanism for non-LTR retrotransposition". Cell. 72 (4): 595–605. doi:10.1016/0092-8674(93)90078-5. PMID 7679954.
- Christensen, SM; Ye, J; Eickbush, TH (2006). "RNA from the 5′ end of the R2 retrotransposon controls R2 protein binding to and cleavage of its DNA target site". Proceedings of the National Academy of Sciences of the United States of America. 103 (47): 17602–17607. Bibcode:2006PNAS..10317602C. doi:10.1073/pnas.0605476103. PMC 1693793. PMID 17105809.
- Ruschak AM, Mathews DH, Bibillo A, et al. (2004). "Secondary structure models of the 3′ untranslated regions of diverse R2 RNAs". RNA. 10 (6): 978–987. doi:10.1261/rna.5216204. PMC 1370589. PMID 15146081.
- Eickbush, DG; Eickbush, TH (Jul 2010). "R2 Retrotransposons Encode a Self-Cleaving Ribozyme for Processing from an rRNA Cotranscript". Molecular and Cellular Biology. 30 (13): 3142–3150. doi:10.1128/MCB.00300-10. PMC 2897577. PMID 20421411.
- Kierzek, E., Kierzek, R., Moss, W. N., Christensen, S. M., Eickbush, T. H. & Turner, D. H. (2008). Isoenergetic penta- and hexanucleotide microarray probing and chemical mapping provide a secondary structure model for an RNA element orchestrating R2 retrotransposon protein function. Nucleic Acids Res 36, 1770-82.
- Kierzek E., Christensen S.M., Eickbush T.H., Kierzek R., Turner D.H., Moss W.N. (2009). Secondary structures for 5’ regions of R2 retrotransposon RNAs reveal a novel conserved pseudoknot and regions that evolve under different constraints. J Mol Biol 390: 428–442.
- Moss WN, Eickbush DG, Lopez MJ, Eickbush TH, and Turner DH (2011). The R2 retrotransposon RNA families. RNA Biol 8(5): 714–718.