HIST1H2BB
Histone H2B type 1-B is a protein that in humans is encoded by the HIST1H2BB gene.[5][6][7]
| H2BC3 | |||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
 | |||||||||||||||||||||||||
| |||||||||||||||||||||||||
| Identifiers | |||||||||||||||||||||||||
| Aliases | H2BC3, H2B.1, H2B/f, H2BFF, histone cluster 1, H2bb, histone cluster 1 H2B family member b, HIST1H2BB, H2B clustered histone 3 | ||||||||||||||||||||||||
| External IDs | OMIM: 602803 MGI: 2448377 HomoloGene: 137348 GeneCards: H2BC3 | ||||||||||||||||||||||||
| |||||||||||||||||||||||||
| |||||||||||||||||||||||||
| |||||||||||||||||||||||||
| |||||||||||||||||||||||||
| Orthologs | |||||||||||||||||||||||||
| Species | Human | Mouse | |||||||||||||||||||||||
| Entrez | |||||||||||||||||||||||||
| Ensembl | |||||||||||||||||||||||||
| UniProt | |||||||||||||||||||||||||
| RefSeq (mRNA) | |||||||||||||||||||||||||
| RefSeq (protein) | |||||||||||||||||||||||||
| Location (UCSC) | Chr 6: 26.04 – 26.04 Mb | Chr 13: 23.75 – 23.75 Mb | |||||||||||||||||||||||
| PubMed search | [3] | [4] | |||||||||||||||||||||||
| Wikidata | |||||||||||||||||||||||||
| |||||||||||||||||||||||||
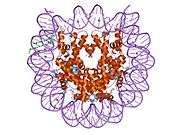
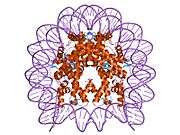
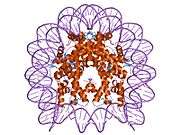
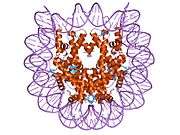
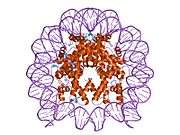
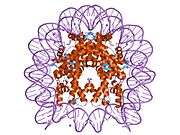
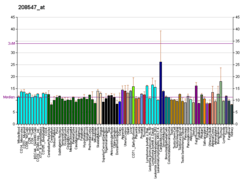
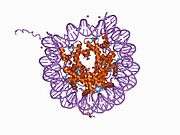
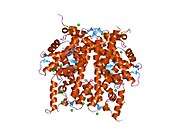
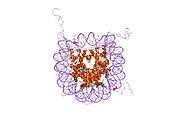
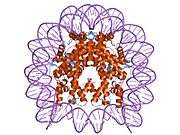
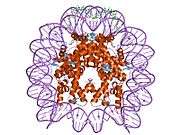
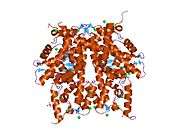
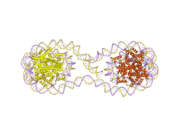
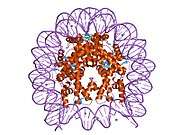
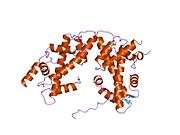
Histones are basic nuclear proteins that are responsible for the nucleosome structure of the chromosomal fiber in eukaryotes. Nucleosomes consist of approximately 146 bp of DNA wrapped around a histone octamer composed of pairs of each of the four core histones (H2A, H2B, H3, and H4). The chromatin fiber is further compacted through the interaction of a linker histone, H1, with the DNA between the nucleosomes to form higher order chromatin structures. This gene is intronless and encodes a member of the histone H2B family. Transcripts from this gene lack polyA tails; instead, they contain a palindromic termination element. This gene is found in the large histone gene cluster on chromosome 6p22-p21.3.[7]
References
- GRCh38: Ensembl release 89: ENSG00000276410 - Ensembl, May 2017
- GRCm38: Ensembl release 89: ENSMUSG00000075031 - Ensembl, May 2017
- "Human PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- "Mouse PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- Kardalinou E, Eick S, Albig W, Doenecke D (Dec 1993). "Association of a human H1 histone gene with an H2A pseudogene and genes encoding H2B.1 and H3.1 histones". J Cell Biochem. 52 (4): 375–83. doi:10.1002/jcb.240520402. PMID 8227173.
- Marzluff WF, Gongidi P, Woods KR, Jin J, Maltais LJ (Oct 2002). "The human and mouse replication-dependent histone genes". Genomics. 80 (5): 487–98. doi:10.1016/S0888-7543(02)96850-3. PMID 12408966.
- "Entrez Gene: HIST1H2BB histone cluster 1, H2bb".
Further reading
- Albig W, Kardalinou E, Drabent B, et al. (1991). "Isolation and characterization of two human H1 histone genes within clusters of core histone genes". Genomics. 10 (4): 940–8. doi:10.1016/0888-7543(91)90183-F. PMID 1916825.
- Albig W, Kioschis P, Poustka A, et al. (1997). "Human histone gene organization: nonregular arrangement within a large cluster". Genomics. 40 (2): 314–22. doi:10.1006/geno.1996.4592. PMID 9119399.
- Albig W, Doenecke D (1998). "The human histone gene cluster at the D6S105 locus". Hum. Genet. 101 (3): 284–94. doi:10.1007/s004390050630. PMID 9439656.
- El Kharroubi A, Piras G, Zensen R, Martin MA (1998). "Transcriptional activation of the integrated chromatin-associated human immunodeficiency virus type 1 promoter". Mol. Cell. Biol. 18 (5): 2535–44. doi:10.1128/mcb.18.5.2535. PMC 110633. PMID 9566873.
- Deng L, de la Fuente C, Fu P, et al. (2001). "Acetylation of HIV-1 Tat by CBP/P300 increases transcription of integrated HIV-1 genome and enhances binding to core histones". Virology. 277 (2): 278–95. doi:10.1006/viro.2000.0593. PMID 11080476.
- Deng L, Wang D, de la Fuente C, et al. (2001). "Enhancement of the p300 HAT activity by HIV-1 Tat on chromatin DNA". Virology. 289 (2): 312–26. doi:10.1006/viro.2001.1129. PMID 11689053.
- Galasinski SC, Louie DF, Gloor KK, et al. (2002). "Global regulation of post-translational modifications on core histones". J. Biol. Chem. 277 (4): 2579–88. doi:10.1074/jbc.M107894200. PMID 11709551.
- Strausberg RL, Feingold EA, Grouse LH, et al. (2003). "Generation and initial analysis of more than 15,000 full-length human and mouse cDNA sequences". Proc. Natl. Acad. Sci. U.S.A. 99 (26): 16899–903. doi:10.1073/pnas.242603899. PMC 139241. PMID 12477932.
- Cheung WL, Ajiro K, Samejima K, et al. (2003). "Apoptotic phosphorylation of histone H2B is mediated by mammalian sterile twenty kinase". Cell. 113 (4): 507–17. doi:10.1016/S0092-8674(03)00355-6. PMID 12757711.
- Lusic M, Marcello A, Cereseto A, Giacca M (2004). "Regulation of HIV-1 gene expression by histone acetylation and factor recruitment at the LTR promoter". EMBO J. 22 (24): 6550–61. doi:10.1093/emboj/cdg631. PMC 291826. PMID 14657027.
- Citterio E, Papait R, Nicassio F, et al. (2004). "Np95 is a histone-binding protein endowed with ubiquitin ligase activity". Mol. Cell. Biol. 24 (6): 2526–35. doi:10.1128/MCB.24.6.2526-2535.2004. PMC 355858. PMID 14993289.
- Gerhard DS, Wagner L, Feingold EA, et al. (2004). "The status, quality, and expansion of the NIH full-length cDNA project: the Mammalian Gene Collection (MGC)". Genome Res. 14 (10B): 2121–7. doi:10.1101/gr.2596504. PMC 528928. PMID 15489334.
- Golebiowski F, Kasprzak KS (2007). "Inhibition of core histones acetylation by carcinogenic nickel(II)". Mol. Cell. Biochem. 279 (1–2): 133–9. doi:10.1007/s11010-005-8285-1. PMID 16283522.
- Zhu B, Zheng Y, Pham AD, et al. (2006). "Monoubiquitination of human histone H2B: the factors involved and their roles in HOX gene regulation". Mol. Cell. 20 (4): 601–11. doi:10.1016/j.molcel.2005.09.025. PMID 16307923.
- Bonenfant D, Coulot M, Towbin H, et al. (2006). "Characterization of histone H2A and H2B variants and their post-translational modifications by mass spectrometry". Mol. Cell. Proteomics. 5 (3): 541–52. doi:10.1074/mcp.M500288-MCP200. PMID 16319397.
- Pavri R, Zhu B, Li G, et al. (2006). "Histone H2B monoubiquitination functions cooperatively with FACT to regulate elongation by RNA polymerase II". Cell. 125 (4): 703–17. doi:10.1016/j.cell.2006.04.029. PMID 16713563.
- Kim SC, Sprung R, Chen Y, et al. (2006). "Substrate and functional diversity of lysine acetylation revealed by a proteomics survey". Mol. Cell. 23 (4): 607–18. doi:10.1016/j.molcel.2006.06.026. PMID 16916647.