Autophagy database
Autophagy database(s) aim to provide a comprehensive list of autophagy-related genes and proteins, whether they are identified as orthologs or homologs of other, potentially related, proteins. Many kinds of information, including sequences, functions, and 3D structures, can be stored, thus making them accessible in a searchable format. Information available in a single source, using a searchable format, would simplify work for future researchers. These sources would then help to accomplish this aim by providing recently published references on autophagy alongside categories such as user ratings, a list of informative reviews, and results of an original analysis. As autophagy can play a role in a host of human diseases, such as those of the heart, liver, and kidney, further understanding its mechanisms is essential. Simplifying the research process with some database would then provide a scientific boon.[2]
| AUTOPHAGY DATABASES | |
|---|---|
| Content | |
| Description | Databases to categorize autophagy-related genes and proteins |
| Contact | |
| Research center | Center for Information Biology; DNA Data Bank of Japan, Luxembourg Institute of Health |
| Primary citation | Homma et al., [1], Luxembourg Institute of Health [2] |
| Release date | 2010 |
| Access | |
| Website | tp-apg |
| Tools | |
| Web | autophagy |
Autophagy
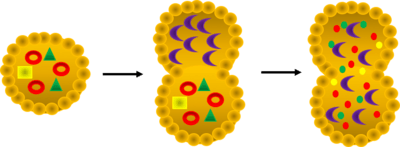
[For a complete background, please refer to Autophagy].
Autophagy is the process by which the cells in an organism destroy non-functional or unnecessary self-components.[3] Specifically, autophagy is a catabolic process involving the degradation of a cell's own components through the lysosomal machinery.[1] Autophagy is also crucial for instances of starvation and removal of potentially dangerous cellular materials, indicating its necessity in maintaining life.[1] As seen in the associated figure Autophagy, cellular products are degraded by destructive cellular components, such as lysosomes, to produce new materials for the cell to use. Research into autophagy and its related processes has exploded over recent years, however, many of these processes are not completely understood and homologs have not been found in different species for many of these proteins.[1] Its molecular mechanisms have not been fully elucidated, despite dramatic advances in the field as evidenced by hundreds of autophagy-related genes and proteins reported.[1] As such, there was a demonstrated need for a database to characterize human autophagy proteins and components and/or their homologs, as well as orthologs in other species.
Autophagy database
Autophagy database is a product of the National Institute of Genetics (NIG) [4] NIG was founded in June 1949 by the ministry of Education, Science, Sports, and Culture, with Prof. Kan Oguma being elected the first director.[4] Over time, many departments have been added for various applications such as Genetics, Genomics, DNA Research, and, most notably for our purposes, the DNA Data Bank.[4] NIG is a division of the Japanese Research Organization of Information and Systems, and is currently under the supervision of its ninth director.[4] NIG aims to conduct top-level research in the pursuit of streamlining of information, as well as the dissemination of information from research into societal application.[4] A tool created by this organization for this purpose is the Autophagy database.
The Autophagy database is a database of proteins involved in autophagy. The Autophagy database intends to collect all relevant information, organize it, and make it publicly available so that its users can easily get up-to-date knowledge. Specifically, the Autophagy database offers a "free-for-all" tool for those with interests, research and otherwise, in autophagy.[3] To better accomplish this aim, the available Autophagy database from NIG calls for users of the database to disseminate and share information, so that autophagy-related data can be available for free to all who need it.[1] For an interested research community, this model of research dissemination holds promise. As of April 2018, there were 582 reviewed proteins available in this database.[3] Including autophagic proteins available in HomoloGene, NCBI, there are over 52,000 total proteins.[3] Autophagy database offers comparison of homologous proteins between 41 different species to search new and old autophagy-related proteins, so that current autophagy research can be streamlined.[1] The database was made publicly available in March 2010 and currently includes 7,444 genes/proteins in 82 eukaryotes.
Human autophagy database
Human autophagy database is a product of the Luxembourg Institute of Health (LIH).[5] LIH has several branches throughout Luxembourg available for Biomonitoring, Infection and Immunity, Health administration, Oncology, Sports Medicine, and Biobank. Each of these departments aims to support the LIH mission statement, which is "to generate and translate research knowledge into clinical applications with an impact on the future challenges of health care and personalised medicine."[5] It offered tools.[5] The Laboratory of Experimental Cancer Research of LIH helped to establish one of these tools, that tool being the database known as Human autophagy database.
Human autophagy database (HADb) is another available autophagy resource. [2] Unlike Autophagy database, Human autophagy database only compares those proteins found in humans. HADb is the first human-only autophagy database, where researchers may find an updated listing of directly and indirectly related autophagic proteins, given no consistent database previously available to compensate for a huge expansion in autophagy research.[2] HADb does not only provide information on the gene of interest, but also aims to evolve into a database which can be used to analyze the gene of interest.[2] For this purpose, HADb was made as complete as possible in terms of autophagy-related proteins, though newly discovered proteins and genes may be submitted by different users to the Submission section. The information provided by Human autophagy database can be used further in bioinformatics applications.
Use in bioinformatics
Given that these databases are a large store of biological information, these can be used in bioinformatics applications to simplify information collection and analysis. Bioinformatics looks to pair biological discoveries with big data, to aid in improved scientific discoveries. Each database can be utilized to study an autophagic protein or gene of interest, where these databases are maintained by user submissions. Information for each gene can be used to access Entrez, Ensembl, and PubMed. FASTA sequence is also available for sequence analysis using sites such as BLAST. Specific uses available to Autophagy database and Human autophagy database are shown below.
Autophagy database has several available functions to search for autophagy-related proteins in different species.
A user may access Autophagy database at http://www.tanpaku.org/autophagy/index.html. The image given, "Options for ADb", showcases the variety of options available for this database. All unhighlighted tabs offer additional information and contact information unrelated to gene search. A user may refer to:

- the Protein list, highlighted yellow, where the user may select an organism and search for Synonyms, Gene ID, and Protein accession, among other functions. These function offer the user multiple options on how to search for information on genes of interest. The options available on Autophagy database for Protein list can be seen in the example given to the right Protein list given to the right. Selecting various options, such as Synonyms, allows the user to search using specific queries.
- matches of autophagy-related proteins amongst homologs and orthologs under the Homologs tab, highlighted in green. This can be used according to the image to the right, Homologs, where orthologs and homologs can be compared between different organisms and taxa by selecting the required boxes.
- search for specific genes. This may be accomplished using Keyword search, highlighted in blue, and may also be used to match a known gene of interest to a certain species. This helps to determine potential orthologs.
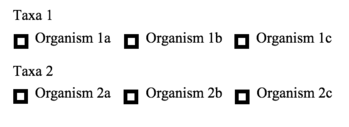
- homologs and orthologs to a gene of interest. Homology search, highlighted in orange, can be used to search a FASTA or unformatted text sequence of a known gene. This may aid in finding connected autophagy-related proteins, or in finding homologs or orthologs. Homologs and orthologs can be compared by the user within or amongst species, given different species and taxa options as seen in the associated figure.
- analyze connections between one gene and autophagy genes available in the database. Original Analyses, highlighted in grey, may be used to find potential autophagy-related gene matches to a known gene. To best utilize these functions, the user should refer to the "Download" tab to download all gene files, so that function of Autophagy database can be fully utilized.
Human autophagy database has available functions for Look for gene and Clustering.[2]
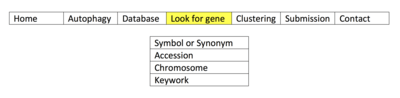

When accessing http://autophagy.lu/index.html, these options can be accessed. Interested parties may also Submit new human autophagy proteins to the database. A user may utilize the database according to the following options:
- A user may search for a gene by name in Look for gene category. A gene may be sought for using its gene symbol, Ensembl accession number, chromosome location, or a relevant keyword. Simple instructions for how to access a gene of interest using this method are given in the figure for "Look for gene: HADb". Briefly, the user would access the website, select the highlighted tab, and search for their gene of interest using the available tabs given in the associated image. Specifically, the user would select which option they would like to use in the associated table, and then fill in the information for the desired tab, whether it be Symbol or Synonym, Chromosome, Accession number, or Keyword. Once the tab is selected, information can be entered and searched to determine any linked autophagy-related proteins.
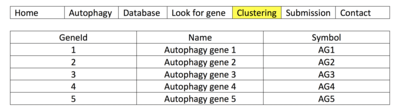
- The user may also refer to Clustering where genes may be viewed in alphabetical order. A simplified map of how to conduct this search is shown in image "Clustering: HADb". The user would first refer to http://autophagy.lu/index.html, after which they would select the highlighted tab in the "Clustering" image to access their gene of interest. The user can then select their gene of interest alphabetically to gather further information. Though this database contains only human autophagy genes, the user need not download a database for use, and can find genes and proteins involved in the complex process of autophagy.
Limitations of the Databases
Each database offers its own strengths and weaknesses.
- Autophagy database: Conceptually, Autophagy database offers the opportunity to easily access information on autophagy-related proteins in a variety of species.[3] However, there are some issues in using this database. The user may try the assortment of tab options available Options for ADb, though these options cannot be utilized without downloading content from Autophagy database. Though the user can access the Download tab (seen in "Options for ADb" in white), this offers only text output when using U.S.-based wifi service. As such, the user cannot access the variety of options mentioned above in a GUI format, but rather must search text output. The user may download the Autophagy database, but these files may be difficult to access using Apple OS. This complicates the ease of use for the U.S.-based user, reducing Autophagy database's utility. This is a potential complication of an internationally available database, that complicates its ease of use.
- Human autophagy database: Though also an internationally available database, ease of use for Human autophagy database is considerably improved. All available options, though limited, can be accessed using a U.S.-based wifi service. Human autophagy database is limited in the array of options available for data collection and analysis, as there are fewer options available than those offered by Autophagy database. The database also stores only human autophagy-related genes and proteins,[2] whereas Autophagy database has information on autophagy-related genes and proteins available for a variety of different species.
Benefits of the Databases
Though each database has its own strengths and weaknesses, they each help to fill a gap.[3] Further additions may help to improve these databases in the future. Though there may be databases available that appear more complete for general gene or protein searches, such as NCBI, HADb and Autophagy database offer the most complete information on autophagy-related genes and proteins. The GUI is not fully refined for each, and may be harder to access, but each of these databases maintains focus on autophagy, whereas NCBI does not use the same focused approach on autophagy. As such, HADb and Autophagy database may offer an interesting route for exploration of autophagy-related genes and proteins.
See also
References
- Homma, Keiichi; Suzuki Koji; Sugawara Hideaki (Jan 2011). "The Autophagy Database: an all-inclusive information resource on autophagy that provides nourishment for research". Nucleic Acids Res. England. 39 (Database issue): D986-90. doi:10.1093/nar/gkq995. PMC 3013813. PMID 20972215.
- Luxembourg Institute of Health. (n.d.). HADb: Human autophagy database. Retrieved April 13, 2018, from http://autophagy.lu/index.html
- Homma, K., Suzuki, K., & Sugawara, H. (n.d.). Autophagy database. Retrieved April 13, 2018, from http://www.tanpaku.org/autophagy/index.html
- National Institute of Genetics. (2018). National Institute of Genetics. Retrieved April 23, 2018, from https://www.nig.ac.jp/nig/
- Luxembourg Institute of Health. (2015). Luxembourg Institute of Health. Retrieved April 23, 2018, from https://www.lih.lu/
External links
- https://web.archive.org/web/20110309173256/http://tp-apg.genes.nig.ac.jp/autophagy/
- Another database specializing in human autophagy (http://autophagy.lu/) is also publicly available.