22
4
A little genetics lesson
When you only have access to someone's visible traits or phenotype, a pedigree of their family history is often used to figure out the actual genetic information or, genotype of each family member.
When dealing with simple dominance as we will be, a simple pedigree chart will be enough to figure out the alleles, or the version of the genes that they have, of each person. In simple dominance a person with a dominant allele (denoted with a capital letter) will always have the trait that that version represents, no matter the other allele. It takes two recessive alleles (denoted with a lowercase letter) to have that version expressed. In other words the dominant allele always masks the recessive version of that gene. Here is an example of a pedigree chart:
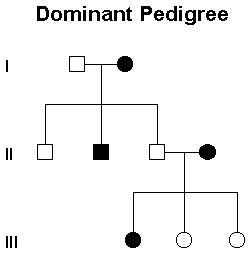
Each row here is a generation. Circles are female, males squares, horizontal lines are marriage, vertical lines children. Pretty simple. Black means recessive phenotype, white, dominant. Starting from the top, (assume the alleles are A and a), we know person 2 has aa, homozygous recessive because that is the only possible option for the recessive phenotype. Now even though person one could be either Aa or AA to be dominant phenotype, because he has a recessive child, he must be Aa, or heterozygous. You can do this for all the other people. In the event you don't have any information that enables you to figure out the second allele, it can be done like so: A_.
Your Task
- You will receive a pedigree chart in the form of a list of generations like
[GenI, GenII, etc.]in any sane format. - Each generation will be a list of strings, each string representing a person.
- The people are made up of three parts - an ID, their phenotype, and their "connections".
- Their ID is a single printable ascii character that is unique in the entire tree other than
Aora. (No, there won't be more than 95 people in the chart). - Their phenotype is one of
Aora,Abeing the dominant allele, andabeing recessive. - Their connections are a sequence of ID's of other people that they have connections with.
- A connection in the same generation is marriage, in different generations is child and parent.
- The connections are repeated on both sides (i.e. the husband has say he is a husband of wife, and wife says she is husband of wife).
- You have to figure out the genotypes of everyone as much as possible.
- Return the same list, except instead of people, put their genotypes in the same position.
- The genotype has to be outputted in order so
Aainstead ofaA. - A little leeway on the input format is fine.
- This is code-golf so shortest answer in bytes wins.
Examples
[["0A1234", "1a0234"], ["2A01", "3a01", "4A015678",
"5a4678"], ["6a45", "7A45","8A45"]] (The one above) ->
[["Aa", "aa"], ["Aa", "aa", "Aa", "aa"], ["aa", "Aa", "Aa"]]
[["0A12", "1A02"], ["2A301", "3a2"]] ->
[["A_", "A_"], ["A_", "aa"]]
Bonus
- -30 bytes if you deal with incomplete and co-dominance also. On the detection of three phenotypes instead of two in the entire chart, apply incomplete/co dominance to your algorithm.
Are we allowed to only modify the
Aandaand leave the ids and connections as is (i.e.[["0A12","1A02"],["2A301","3a2"]]becomes[["0A_12","1A_02"],["2A_301","3aa2"]]instead of[["A_","A_"],["A_","aa"]])? – Kevin Cruijssen – 2018-02-28T14:59:25.783